AbstractThe estuarine ecosystem is under threat due to the addition of different anthropogenic pollutants. Among the various pollutants, microplastics(MPs), and antibiotics play a significant role in affecting estuarine organisms and human health by transport through the food chain. In the estuaries, microorganisms, including pathogens, colonise microplastics by the development of biofilms. Estuaries have long been home to MPs, served as novel hubs for the transmission of Antibiotic Resistance Genes (ARGs). As a result of their continual interactions with a variety of aquatic creatures, MP-associated bacterial communities will eventually present a transfer opportunity to organisms that consume MPs. To comprehend the presence of microplastics and ARGs in the estuaries, a thorough review is necessary. This review discusses the various sources of MPs and antibiotic pollution, as well as the transport of ARGs via biofilm formed on MPs. Furthermore, the factors affecting biofilm formation in estuaries are reviewed. In addition, the transport of ARGs by microbial populations within biofilm is discussed. The ARGs are transported into the food chain, which will be a threat to human health. Hence, the occurrence of antibiotic-resistant genes in fish is discussed.
Graphical Abstract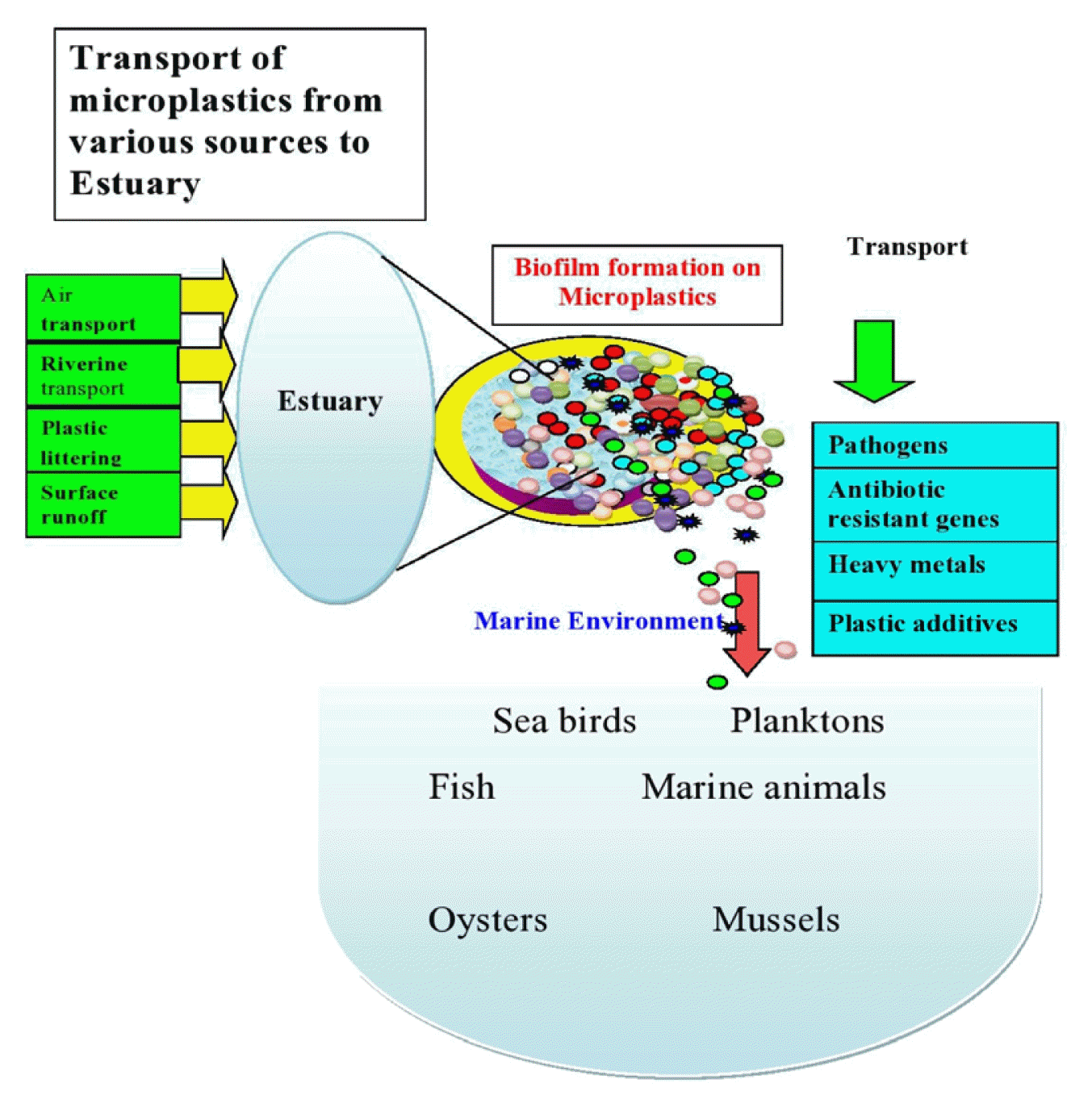
1. IntroductionEstuaries are highly productive and dynamic areas at the transition between rivers and the marine environment. Tributaries transport nutrients into estuaries and transform them into highly productive ecosystems [1,2]. Further, estuaries are vital resources for providing food and habitat for fish and shellfish, food for human consumption, areas for tourism and recreation, and a host of ecosystem services [3–5]. Currently, the estuarine ecosystem is highly polluted [2] by several point and nonpoint sources, including agricultural runoff, wildlife excrement, septic tanks, and sewage discharges [6,7]. Because estuaries are coastal transition zones between freshwater and seawater, pH, temperature, geochemical condition, and salinity vary on a relatively short spatial and temporal scale [8]. The geomorphological and physical-chemical characteristics of estuaries favour the deposition of organic matter and man-made contaminants [9]. The prevalence of low energy and currents from the waves in estuaries makes the pollutants accumulate in sediments instead of dispersing [10]. The types of pollutants found in estuaries are heavy metals [11, 12], radionuclides [13], and pathogens such as bacterial pathogens [14], protozoa [15], enteric viruses [16], and fungus [14]. Furthermore, antibiotics, antibiotic resistant genes, and MPs are regarded as important emergent pollutants in the estuarine ecosystem due to their widespread distribution, persistence, and impact on the environment and human health [17].
Estuaries also play an important role in the transport of pollutants from land to sea [15]. According to research, contaminants are moved from estuaries to the oceans through biofilm that develops on substrates such as MPs [18]. MPs can adsorb toxic pollutants such as antibiotics and heavy metals, both of which have been related to antibiotic resistance [19]. MPs could thus act as vectors for pathogens and antibiotic resistance genes in the aquatic environment [19]. MPs release absorbed toxic pollutants in the marine environment [20] and are consumed by marine biota, including seafood species [21]. Pollutants are indeed being transported into the food chain, which has emerged as a major issue in this decade. As a result, the primary goals of this review are to emphasise the following to highlight the current state, knowledge gaps, and future research directions: (1) the presence of MPs, antibiotics, and ARGs in estuaries; (2) the formation of biofilm on MPs in estuaries; (3) factors influencing biofilm formation on MPs; and (4) the transport of ARGs within the biofilm microbial community.
2. Antibiotic Pollution in EstuariesAntibiotics are used to treat bacterial infections, namely urinary tract infections, sepsis, sexually transmitted diseases, and diarrhea [22]. It is observed that a high rate of resistance has been developed to the antibiotics used to treat these diseases [23]. Antibiotic resistance is accelerated by the overuse or misuse of antibiotics by people [24]. Antibiotic and ARG prevalence is being investigated in agricultural soils [25], rivers [26], and oceans [27]. Besides that, antibiotics pollute the estuaries [28]. Wastewater treatment plants (WWTPs) [29], leachate treatment plants [30], animal husbandry facilities [31], organic dairy farms [32] aquaculture industry [33], poultry farms [34], sewage contamination [35], household and hospital wastewater [36], and biology laboratories [37] are major sources of antibiotics in estuaries. Because antibiotics are not completely metabolised by the human or animal body, functional compounds can enter the aquatic ecosystem via the waste products of humans or animals that have consumed the antibiotics [38]. Furthermore, the types and concentrations of antibiotics in aquatic systems vary globally depending on antibiotic consumption and use [39]. Furthermore, the chemical stability and partition coefficient of antibiotics influence their presence in the aquatic environment [40]. The concentration of antibiotics is very low, and the design of water treatment plants is insufficient to remove the antibiotics [41]. As a result, even treated effluents from WWTPs can be a source of antibiotics in the aquatic environment [42].
Antibiotic resistance is a global health concern; infections with resistant pathogens kill at least 700,000 people each year [43]. According to WHO (2018), pathogenic bacteria resistant to common antibiotics are found in human and veterinary systems. There is an increase in global annual deaths due to antibiotic resistance, which could be increased to 10 million by 2050 [44]. Antibiotic pollution has been found to be more severe in the spring, summer, and winter seasons than in the autumn [45]. In general, antibiotic concentrations in sediment samples were higher than in water samples [46]. Many laboratory scale studies have proven that the presence of antibiotics at concentrations as low as nanograms per millilitre has been shown to promote the development of resistance [47]. Due to the harmful effects of low levels of antibiotics in the environment, antibiotics are now considered an important environmental pollutant [48]. The study conducted in the Yantze river estuary; China showed that the concentrations of antibiotics varied with location [49]. In addition, the highest antibiotic concentration was observed around river discharge and sewage outfall during the January month. Furthermore, there was a positive correlation between total antibiotic concentration and dissolved organic carbon (DOC) concentration [49]. The prevalence of the highest concentration in sediment samples might be due to low flow conditions and low temperatures [50]. The antibiotics, namely tetracycline, sulfaquinoxaline, enrofloxacin, and thiamphenicol, were examined in the Yangtze Estuary and nearby coastal areas [50]. There was a positive correlation between quinolone concentration and Total Organic Carbon (TOC), indicating the significant role played by TOC for the increased accumulation of antibiotics in sediments [50]. In the Jiulong River Estuary, antibiotics such as sulfonamide, quinolon, and chloramphenicol were examined [51]. In India, a study conducted in the Cochin Estuary revealed the occurrence of Vibrio parahaemolyticus which was found to be resistant to antibiotics, namely enrofloxacin, nitrofurantoin, trimethoprim, sulphamethoxazole, streptomycin, amikacin, and nalidixic acid indicating the presence of aforementioned drugs [52]. Further, it was evident that the antibiotic resistant genes were present in plasmids [52]. The abundance of salfonamide-resistant E. coli was reported in Liaohe and Daliaohe Estuaries in China, suggesting the population was higher during the dry season when compared to the wet season [53]. Furthermore, there was significant attenuation of antibiotic-resistant E. coli during movement between land and sea, explaining the role of the estuary as a natural attenuator in reducing the ARGs during the transport of antibiotic-resistant bacteria from rivers to the ocean [53]. In general, ciprofloxacin increased ARG types in biofilm and induced the growth of dominant bacteria belonging to phyla such as proteobacteria, actinobacteria, and planctomycetes [54].
3. Microplastic Pollution in EstuariesMicroplastic pollution is a serious threat to the environment and is considered an emerging contaminant of concern. MPs are heterogeneously mixed plastics that are less than 5 mm in diameter. MPs are distributed in different forms, namely plastic fibres, granules, and pellets [55]. The distribution of MPs is observed from the poles to the equator [56]. Plastics enter the environment primarily as a result of human activity [57, 58]. The important sources of plastic pollution are agricultural mulching [59], sewage sludge disposal [60], wastewater treatment plants [61], informal landfills [62], and biowaste compost [63]. MPs are found in a variety of terrestrial environments, including home and garden soils [64], flood plain soils [65], remote high mountain soils [66], and industrial soils [67], as well as surface and deep ocean waters [68], lakes [69,70], mangroves [71], shorelines [72], and estuaries [73–75]. The studies conducted to examine the distribution of different sizes, forms, and types of MPs in estuaries across the world are presented in Table 1.
According to the investigations, estuaries were filled with MPs in the form of fibre, fragment, foam, film, and pellets [89–93]. Unlike in surface water, MPs from sediment samples were predominantly in the 0.05–0.3 mm size range, making them easily ingestible by aquatic organisms [86]. The shallow sediment contained more microfibers, and the rate of microfiber deposition in the sediment was strongly related to the expected change in production in the study area based on annual plastic production [76]. Furthermore, seawater inundation in the estuary and the distance of sampling points from the coast had a significant influence on the distribution of fibres and films in estuary sediments [77]. The most common polymer types found there were PE, PP, PVC, PA, PAM, and PES [96–98]. However, [78] suggested that MPs carried by upstream river water to the estuary were an important source of MP pollution, and the abundance of MPs varies greatly across regions due to differences in human activities as well as hydrological and hydrodynamic conditions. However, the studies that have been done to look at how different kinds and forms of MPs are distributed in estuaries are insufficient[99]. Because of the vegetation and subsequent flocculation, estuaries can filter pollutants more effectively [100]. Many estuary components, such as coastal wetlands like mangroves, salt marshes, lagoons, and tidal and subtidal flats, act as sediment and pollutant sinks [101]. Because of the extensive weathering nature of microplastics in the environment, their virgin nature is altered, and thus their physiochemical characteristics are altered [102]. Furthermore, changes in crystallinity, specific surface area, and oxygen functional groups cause the sorption behaviour of MPs to change [103]. Furthermore, degradation can be confirmed by a loss of gloss and/or an increase in haze caused by oxidation and cleavage of polymer chains[104]. Semicrystalline polymers, such as PE, experience an increase in crystallinity (and brittleness) as a result of chain scissions in amorphous regions [105]. Photooxidation, on the other hand, causes cracks and increases fragmentation [106].
4. Biofilm Development in EstuariesIn the aquatic environment, bacterial phenotypic plasticity is responsible for bacterial adaptation to different environmental conditions [107,108] which is achieved through the formation of multi-species biofilms, which enables interspecies interactions responsible for ecological processes [109]. Biofilm is commonly composed of microbial cells and their metabolic products, such as extracellular polymers and inorganic precipitates [110]. Bacterial biofilm is thought to be an emerging form of bacterial life in which communal life differs significantly from bacteria that live as free-living cells [111].
Biofilm protects microbes and allows them to survive and thrive in harsh environments. Furthermore, microbes in the biofilm are less susceptible to antibiotics and disinfectants when exposed to them [112]. Bacteria grown in biofilms, on the other hand, can form intricate and complex structures, such as channels that allow nutrients to circulate. The structure of biofilm influences gene expression [113]. Biofilm formation occurs on a variety of substrates, including living tissues, indwelling medical devices, piping in industrial or potable water systems, and natural aquatic systems. Marques et al. [114] investigated the formation of biofilms on hydrophilic surfaces such as stainless steel and glass. The bacterial community in estuaries is dependent on precipitation, according to a study conducted in the Bilbao estuary [2]. Freshwater bacteria from the Comamonadaceae and Spingobacterioaceae were abundant during high precipitation periods. During low precipitation periods, however, marine bacteria from the Rhodobacterales and Oceanospirillales predominated [2].
4.1. MPs as Carriers of ARGsEstuaries, serving as links between land and ocean, harbour numerous microbes that are relatively highly active because of the massive terrigenous input of nutrients [115]. Furthermore, ARGs such as tetracycline, sulfonamides, AmpC β-lactamase, and the class-1 integron gene were found in the leachate treatment plant effluent [116]. ARGs such as sul1, int1, and sul2 were prevalent in the receiving aquatic system. Furthermore, research has found a relationship between ARG concentration and heavy metal concentration [117]. It was observed that the genes encoding antibiotic resistance and heavy metals were found to be located in the same position on chromosomes or mobile genetic elements, which could facilitate the acquisition of multiple resistance genes by bacteria [118]. There was a strong positive correlation between vanadium and gene tetM [119]. Vanadium has physiologically negative effects on a variety of microbes [120], and it can cause bacterial resistance to vanadium via efflux pump tricarboxylic acid cycle enzymes and accelerate horizontal gene transfer [120]. As a result, it is intriguing to speculate that this resistance could confer resistance to ARGs in bacterial communities in the aquatic environment [119]. Metal absorption would also be aided by interactions between metal ions and charged or polar regions of the MP surface, or by non-specific interactions between neutral metal-ionic complexes and the hydrophobic surface of MPs [121].
Since plastic has a longer half-life when compared to natural materials, the plastispheres or microporous structures formed on microplastics provide a favourable condition for attachment and subsequent formation of biofilm by microorganisms [122]. It is evident from the studies conducted so far that weathered polystyrene foams are acting as carriers of antibiotics in the environment [117].
Salmonella sp. and E. coli were the bacterial species that had the ARGs AmpCo, blan-TEM-1, bla-CTX-M, VIM-1, NDM-1, florR, tetG, and mcr-1 [123]. Additionally, the penetration into the biofilms is determined by the type of antibiotic [124]. In contrast to ciprofloxacin, which readily entered the biofilm created by wild-type Klebsiella pneuminiae, ampicillin did not [124]. The major enzymes involved in nitrogen cycling (the anammox, denitrification, and nitrification processes) are altered by antibiotics in aquatic systems, which has an impact on ecosystem performance [125]. Other strains could quickly acquire the plasmid-encoded virulent genes and antibiotic-resistant genes [126]. Table 2 provides a summary of the studies done to determine the occurrence of ARGs in estuaries. Additionally, there have been relatively few studies that look into the sources, characteristics, and distribution of ARGs globally (Table 3). The most common antibiotic resistance genes found in estuaries were Sul, Tet, and Qnr [127, 128]. The antibiotic resistance found in the bacterial isolates, combined with the longevity of plastics in the aquatic environment, suggests that biofilm on plastics has the potential to harbour pathogens and facilitate horizontal gene transfer [129]. The studies show that more research is needed to investigate the distribution of different types of ARGs in estuaries. This could allow us to take appropriate steps to prevent the spread of ARGs into estuaries.
4.2. Biofilm Development on MPs in EstuariesThere are more than 1200 estuaries in more than 120 countries and territories worldwide, including lagoons and fjords [152]. However, the research on the development of biofilms on microplastics was only done in a few numbers of estuaries around the world. Plastic biofilms are an important site for potential pathogen colonisation and horizontal gene transfer since they are long-lasting [129]. The studies carried out to look at the presence of biofilm on MPs in various estuaries around the world are shown in Fig. 1.
The environmental conditions, the bacterial genome, and the components embedded in the biofilm matrix all influence the structure and components of biofilm [153]. The optimal temperature for Salmonella sp. biofilm formation was found to be 25 to 42°C and pH values of 7–8 [154]. A carbon source is also required for biofilm formation [154] Furthermore, the mechanisms involved in biofilm formation differ depending on the bacterial strain and environmental conditions [155]. Surface chemistry and surface energy are vital properties for microorganism attachment. Furthermore, four distinct three-dimensional architectures were observed when immersing stainless steel and polycarbonate surfaces in a biofilm in an estuary [156]. For example, mushroom like structures, homogeneous, hairy biofilms with horizontal fibres and heterogeneous tousled biofilms were formed by Flavobacterium sp. 112003, Roseobacter sp. IV3009, Shewanella sp. IV3014 and Roseovarius sp. VA04, respectively [156]. The antibiotic resistance detected among the isolates coupled with longevity of plastics in the aquatic environment suggests biofilm on plastics have potential to pathogens and horizontal gene transfer [129]. The bacterial community structure of biofilm developed on different types of MPs in estuaries is shown in Table 4.
It is evident from previous studies that MPs can readily release hazardous pollutants like DDT, polybrominated diphenyl ethers, and other additives into the aquatic environment, thereby elevating the concentration [166]. Also, MPs are transporting organic pollutants into the environment [167]. In the biofilm, a variety of microorganisms with varying redox potential requirements occur in close proximity [168]. In general, preferential attachment of cells during biofilm formation is achieved by the different topography of the substrates, namely, irregular surfaces, etching surfaces, and grooved surfaces [169], when compared to flat surfaces. Furthermore, it is clear that biofilm formation is strongly influenced by environmental nutrient concentrations, such as nitrogen [170] and carbon sources (mannose and trehalose) [171]. In general, sessile microorganisms in biofilms have high environmental stability, more opportunities for interactions such as horizontal gene transfer and co-metabolism are resistant to physical disturbances and antimicrobial compounds [172]. During biofilm formation, the initiation of corrosion reactions by microorganisms is achieved by the formation of concentrated cells, the production of corrosive metabolites, the dissolution of protective layers, the uptake of electrons directly from the substrate, and the production of non-protective layers [173]. In addition, the high population densities, and proximity of cells in biofilms increase the chances for genetic exchange among the bacterial species, converting biofilms into hot spots of antibiotic resistance [174].
4.3. Factors affecting MP associated Biofilm FormationThe important factors affecting MP associated biofilms are size and shape of MPs, hydrophobicity, plastic additives, presence of phenolic compounds and heavy metals [162, 175, 176]. The different factors affecting the biofilm formation on MPs and the effect of biofilm on MPs is illustrated in Fig. 2.
Few studies demonstrated that the microbial community structure of biofilm formed on MPs is influenced by polymer type. Debroas et al. [177] examined that PS, PE, and PET plastic particles were dominated by Alphaproteobacteria and Gammaproteobacteria while Burkholderiales dominated on PE MP biofilms. Further, the early colonizers on PE, and PS MPs were found to be Vibrio sp. [178]. However, the dominant PE colonizing microbial community varies during biofilm formation [129]. The biofilm structure and function depend on microbial growth rate and the type and level of energy sources [179]. In general, microorganisms colonized in the upper regions of the biofilm are more active due to greater accessibility to nutrients and capable of discharging metabolic waste products compared to cells at the bottom region [180]. The biofilm developed on three types of MPs prevalent in the environment namely HDPE, PET, and PS was examined. The virgin MPs were incubated for a period of 31 days with water collected along freshwater - estuarine gradient of the Raritan River in New Jersey, USA. The study revealed that salinity and polymer type had impact on formation of biofilm in which colonization of distinct microbial community was achieved. One of the most abundant MP colonizing bacterial species was found to be Limnobacter thiooxidans [181].
The quantum of chemical energy possessed by plastics is greater, but it is unclear how biofilm formation and microbes facilitate the degradation of plastic debris [182]. There is a relationship between biofilm formation and the physicochemical characteristics of polymers. The characteristics of the plastics, namely compression, crystallinity, surface chemistry, hydrophobicity, and surface topography of plastic particles, determine the microbial community structure in the biofilm [183]. Spingobium, Novospingobium, and uncultured Planctomycetae were the predominant microbial types colonised on polyethylene MPs [183]. Further, significant changes in crystallinity of PE, stiffness of PP, and compression of PS were observed due to biofilm formation [183]. The PE degrading strain of the actinomycete Rhodococcus ruber showed laccase activity and reduced the molecular weight of PE [184]. However, the biofilm formed in the LDPE-Titania environment increased hydrophobicity by the different bacterial species, namely Pseudomonas aeruginosa (CA9), Burkholderia seminalis (CB12), and Stenotrophomonas pavanii (CC18) [185]. The factors influencing the colonisation of microorganisms in biofilms developed on MPs are summarised in Table 5.
Plastics are degraded by the microbial community colonised by plastic particles [185]. Microorganisms achieved preferential degradation of amorphous regions and non-preferential degradation of plastic surfaces in general [192]. Vaksmaa et al. [183] discovered the presence of various bacterial communities on plastics belonging to the orders Flavobacteriales, Alteromonadales, Chitinophagales, and Oceanospirillales. Furthermore, Ideonella sakaiensis [193] degraded PET, Alcanivorax borkumensis [194] degraded low density polyethylene (LDPE), Zalerion maritimum [195] degraded PE, and Aspergillus sp. [196] degraded PE (HDPE). On the surface of the PE MPs, a wide range of biofilm morphological types were observed, including coccus, rod-shaped, and disc-shaped bacterial cells, interwined filaments, and a dense layer of EPS [186]. Similarly, the hydrophobicity of the PE decreased with exposure time. It could be because the surfaces of the MPs absorb organic and inorganic nutrients from water in a matter of hours [186]. This results in the formation of a conditioning film, which can quickly attract microorganisms and facilitate the utilisation of nutrients on said surfaces [186]. The larger the contact angle, the greater the hydrophobicity of the solid surface. It was found that the contact angles for PA, PET, and PVC were 70°, 81°, and 87° respectively. Hydrophyllic surfaces are typically associated with contact angles less than 90° [188]. The PVC with the lowest negative ζ-potential showed the highest structural diversity after incubation in seawater, indicating that the structural diversity is additionally dependent on the incubation medium [188]. Temperature had an impact on the formation of a biofilm on MPs, which was connected to metabolic activity and enzyme activity [175].
4.4. MPs as Carrier of Antibiotics and ARGsWhen compared to virgin materials, biofilm formed on MPs can absorb more pollutants. Furthermore, MPs in biofilm can transport pollutants and microorganisms to aquatic ecosystems [167,197]. Besides that, the toxic effect of plastics can be determined by particle size and surface modification. Nanoplastic-induced oxidative stress can reduce biofilm enzyme activity [176]. Studies confirmed that the surface properties of LDPE microplastics influenced the biofilm communities that formed on them. However, pathogens were more prevalent in biofilm colonised on LDPE microplastics [198].
The occurrence of antibiotic resistance genes was studied by researchers in estuaries. Biofilms are considered as hot spots of antibiotic resistance [199] due to the exchange of genetic elements among bacterial species, which is achieved by high population density and proximity of cells [174]. When susceptible microorganisms in biofilms are exposed to antibiotics in the environment, they become resistant to antibiotics due to selective pressure, chromosomal mutations, and horizontal gene transfer. It is understood from previous studies that ARG transfer is mainly achieved by horizontal gene transfer [200]. The genes responsible for antibiotic resistance are located on mobile genetic elements present in bacteria, namely plasmids, transposons, and membrane vesicles, integrons and membrane vesicles [201] cause horizontal gene transfer [202]. Some bacteria carry plasmids, which provide multiple antibiotic resistance and allow them to survive in harsh environments [203]. Antibiotic-resistant bacteria that have detached from biofilms can spread in the environment and affect human health [199]. Furthermore, antibiotics and ARGs influence nutrient cycling in aquatic systems by altering bacterial community related to biogeochemical cycling Zeng et al. [204]. The sources of ARGs and horizontal gene transfer between microorganisms colonized within biofilm developed on MPs are shown in Fig. 3.
The mobility of ARG transfer between cell donor and recipient is achieved by conjugation, transformation, and transduction. However, intracellular transport is facilitated by integrons and transposons [205]. The study conducted by Ma and Bryers [206] revealed that the bacteria in biofilm sense the antibiotics they are resistant, which enhances the spread of antibiotic resistant. It is suggested that high surface/volume ratio of biofilm favors the efficient transfer of genes between or within the microbial populations [207]. Many studies were conducted to examine the conjugation mediated horizontal transfer of ARGs by bacterial species namely E. coli [145] and Pseudomonas putida [208]. The transformation was another way transfer of genetic element from donor to recipient in Bacillus subtilis and Acinetobacter baylyi ADP1 [209]. In addition, gene transfer is achieved by transduction in which E. coli and Flavobacterium sp. were the donor and recipient respectively [210]. The study conducted by Solheim et al. [211] revealed that phage mediated stx-gene transfer was facilitated between E. coli O103:H25:1199 (recipient) and E. coli C600 731 within biofilm.
MPs are the niche for many microorganisms and provide microhabitats, especially for opportunistic pathogens. In the aquatic systems, MPs selectively enrich ARGs within the biofilm [212]. Interactions between MPs and ARGs are mediated by van der Waals forces, electrostatic forces, and π- π - interactions between MPs and ARGs [212]. It was observed that the bacterial community on MPs possessed higher average path length, clustering coefficient, and modularity compared to those in water samples [167]. The MPs examined in the rivers Jiang and Zhejiang in China possessed a relative abundance of integrin-integrase gene classes 1 and 2, which facilitate horizontal gene transfer [167]. But limited studies have proven that the genetic materials collectively known as resistomes provide molecular function that protects the bacteria against many antibiotics [213]. He et al. [214] demonstrated that tetracycline resistant profiles and ARGs were observed in Listeria spp. isolated from catfish fillets. However, Aeromonas sp. isolated from the gut, liver, skin, mucus, gills, and flesh of rainbow trout harboured aadA, sul1, and tet genes [215]. The ARGs, namely blaDHA, blaACC, tetV, tetM, mdtE, mexF, vanTG, vanWG, vgaB, msrA, and mphA, were examined in fillets of rainbow trout fish, and the antibiotic resistance genes were carried by Carnobacterium sp. and Psedomonas sp. [216]. Miller, Harbottle [217] observed severe mortalities in larvae caused by Vibrio harveyi resistant to trimethoprim-sulfamethoxazole, chloramphenicol, erythromycin, and streptomycin. In addition, Aeromonas sp. colonised in carp harvested from pond water was found to be resistant to β-lactam and lincosamide antibiotics [218].
The antibiotic resistance genes (ARGs), namely tetA, tetO, tetQ, tetW, sulI, sulII, and blaTEM-1, and class 1 integrase (intI1) in the range of 9.4 ×10−6 − 1.6×10−1 and 6.7×10−5 − 5.2×10−1 gene copies per 16S rRNA gene, respectively, were observed in the guts of four major Chinese freshwater carp (i.e., silver carp, grass carp, bighead carp, and crucian carp) from food retail markets. The antibiotic resistance genes were present in Acidobacteria, Bacteroidetes, Chloroflexi, Cyanobacteria, Firmicutes, and Proteobacteria [219]. In addition, V. harveyi isolated from a shrimp farming facility was found to be resistant to ciprofloxacin, penicillin, rifampicin, and vancomycin [220].
5. Conclusions and PerspectivesBased on the preceding discussion, it is possible to conclude that estuaries all over the world are under threat from antibiotics, antibiotic resistant genes (ARGS), and MP pollution. Agricultural soils, animal husbandry facilities, sewage treatment plants, and aquaculture systems are major sources of ARGs and microplastics. MPs found in estuaries are an important substrate for the colonisation of various microorganisms with ARGs. Furthermore, the polymer type influences the microbial community structure in biofilms formed on MPs. Horizontal gene transfer also allows ARGs to be transferred between microorganisms within a biofilm. Furthermore, the prevalence of antibiotic resistance in fish is investigated, resulting in resistance to pathogen-induced infections. Effective wastewater treatment facilities are suggested to prevent the entry of antibiotic, ARG, and MP residues into estuaries. Furthermore, prescribing antimicrobials by human and animal health care professionals based on guidelines will reduce antibiotic overuse, thereby controlling antibiotic accumulation in the environment.
The following study areas could be investigated in light of what is currently known about MP-related biofilm formation in estuaries: (i) The size, colour, and polymer type of MPs are all variable in aquatic environments. MP detection using a single detection technique is challenging. To more accurately detect MPs, multiple detection approaches are needed (ii) There has been little research done to date on the detection of ARGs in aquatic environments like estuaries. If it doesn’t get enough attention, ARGs will move up the food chain and have negative effects. It is crucial to recognise the different ARGs that take place in estuaries. (iii) There isn’t much information currently available to pinpoint the origin of ARGs in estuaries. As a result, source identification is paid greater emphasis, and the appropriate action is required. (iv) It is crucial to think about the bacteria that MPs have colonised. This is accomplished by utilising cutting-edge technologies, like as genomes, metagenomics, proteomics, and metabolomics, to understand more about the formation of biofilms and the biodegradation of MPs. (v) In estuaries, the mechanism of biofilm development on MPs of various sizes and types is poorly understood. It is necessary to pay more attention to how physical, chemical, biological factors, and MP interactions influence biofilm formation and the movement of ARGs and hazardous contaminants through the trophic levels in estuaries. (vi) ARG occurrence in estuaries is still out of control. For the protection of aquatic environments, it is necessary to track their behaviour and develop a risk assessment approach.
AcknowledgmentsThe authors thank the Centre for Environmental Studies, Anna University, Chennai, Tamil Nadu, India, for providing research facilities. The authors would also like to thank the two anonymous reviewers who provided helpful suggestions for revising the manuscript.
NotesReferences1. Herlemann DPR, Labrenz , Jürgens K, et al. Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J. 2011;5:1571–1579.
https://doi.org/10.1038/ismej.2011.41
2. Aguirre M, Abad D, Albaina A, et al. Unraveling the environmental and anthropogenic drivers of bacterial community changes in the Estuary of Bilbao and its tributaries. PLoS One. 2017. 26e0178755.
https://doi.org/10.1371/journal.pone.0178755
3. Armitage AR, Weaver CA, Kominoski JS, et al. Resistance to Hurricane Effects Varies Among Wetland Vegetation Types in the Marsh-Mangrove Ecotone. Estuaries Coasts. 2020;43:960970.
https://doi.org/10.1371/journal.pone.0178755
4. Barbier EB. Estuarine and Coastal Ecosystems as Defense against Flood Damages: An Economic Perspective. Front. Cli. 2020;2:594254.
https://doi.org/10.3389/fclim.2020.594254
5. Chilton D, Hamilton DP, Nagelkerken I, et al. Environmental Flow Requirements of Estuaries: Providing Resilience to Current and Future Climate and Direct Anthropogenic Changes. Front. Environ. Sci. 2020;9:764218.
https://doi.org/10.3389/fenvs.2021.764218
6. Cearreta A, Irabien MJ, Leorri E, et al. Recent Anthropogenic Impacts on the Bilbao Estuary, Northern Spain: Geochemical and Microfaunal Evidence. Estuar. Coast. Shelf Sci. 2000. 504571–592.
https://ui.adsabs.harvard.edu/link_gateway/2000ECSS...50..571C/doi:10.1006/ecss.1999.0582
7. Hitchcock JN, Mitrovic SM. Microplastic pollution in estuaries across a gradient of human impact. Environ. Pollut. 2019;247:457–466.
https://doi.org/10.1016/j.envpol.2019.01.069
8. Elliott M, McLusky DS. The Need for Definitions in Understanding Estuaries. Estuar. Coast. Shelf Sci. 2002;55:815–827.
https://doi.org/10.1006/ecss.2002.1031
9. Du Laing G, Rinklebe J, Vandecasteele B, et al. Trace metal behaviour in estuarine and riverine floodplain soils and sediments: A review. Sci. Total Environ. 2009;407(13)3972–3985.
https://doi.org/10.1016/j.scitotenv.2008.07.025
10. Perina FC, Abessa DMS. Contamination and toxicity in a subtropical Estuarine Protected Area influenced by former mining activities. Ocean Coast. Res. 2020;68:e20313.
http://dx.doi.org/10.1590/s2675-28242020068313
11. Senez-Mello TM, Crapez MAC, Ramose Silva CA, et al. Heavy metals bioconcentration in Crassostrearhizophorae: A site-to-site transplant experiment at the Potengi estuary, Rio Grande do Norte, Brazil. Sci. Rep. 2020;10:246.
https://doi.org/10.1038/s41598-019-57152-w
12. Izegaegbe JI, Vivier L, Mzimela HM. Trace metal contamination in sediment in the Mhlathuze Estuary, Northern KwaZulu-Natal, South Africa: effects on the macrobenthic community. Environ. Monit. Assess. 2020;192:401.
https://doi.org/10.1007%2Fs10661-020-08352-9
13. Bolivar JP, Garcia-Tenorio R, Mas JL. Radioactive impact in sediments from an estuarine system affected by industrial wastes releases. Environ. Int. 2002;27(8)639–645.
https://doi.org/10.1016/s0160-4120(01)00123-4
14. Gu Q, Wang W, Li G, et al. Pollution profile of waterborne bacterial and fungal community in urban Rivers of Pearl River estuary: Microbial safety assessment. J. Freshw. Ecol. 2021;6(1)305–322.
https://doi.org/10.1080/02705060.2021.1975580
15. Shapiro K, Conrad PA, Mazet JAK, et al. Effect of Estuarine Wetland Degradation on Transport of Toxoplasma gondii Surrogates from Land to Sea. Appl. Environ. Microbiol. 2010;76(20)6821–6828.
https://doi.org/10.1128%2FAEM.01435-10
16. Robbins PE, Farkas K, Cooper D. Viral dispersal in the coastal zone: A method to quantify water quality risk. Environ. Int. 2019;126:430–442.
https://doi.org/10.1016/j.envint.2019.02.042
17. Mammo FK, Amoah LD, Gani KM, et al. Microplastics in the environment: Interactions with microbes and chemical contaminants. Sci. Total Environ. 2020;743:140518.
https://doi.org/10.1016/j.scitotenv.2020.140518
18. Richard H, Carpenter EJ, Komada T, et al. Biofilm facilitates metal accumulation onto microplastics in estuarine waters. Sci. Total Environ. 2019;683:600–608.
https://doi.org/10.1016/j.scitotenv.2019.04.331
19. Koch N, Islam NF, Sonowal S, et al. Environmental antibiotics and resistance genes as emerging contaminants: Methods of detection and bioremediation. Curr Res Microb Sci. 2021;2:100027.
https://doi.org/10.1016%2Fj.crmicr.2021.100027
20. Gouin T, Roche N, Lohmann R, et al. A Thermodynamic Approach for Assessing the Environmental Exposure of Chemicals Absorbed to Microplastic. Environ. Sci. Technol. 2011;45(4)1466–1472.
https://doi.org/10.1021/es1032025
21. Smith M, Love dC, Rochman CM, et al. Microplastics in seafood and the implications for human health. Curr. Environ. Health Rep. 2018;5:375–386.
https://doi.org/10.1007/s40572-018-0206-z
22. Krupp K, Madhivanan P. Antibiotic resistance in prevalent bacterial and protozoan sexually transmitted infections. Indian J. Sex Transm. Dis Aids. 2015;36(1)3–8.
https://doi.org/10.4103%2F2589-0557.156680
23. Fair RJ, Tor Y. 2014;Antibiotics and Bacterial Resistance in the 21st Century. Perspective. Medicinal Chem 2014. 6:25–64.
https://doi.org/10.4137%2FPMC.S14459
24. Robbins PE, Farkas K, Cooper D. Viral dispersal in the coastal zone: A method to quantify water quality risk. Environ. Int. 2019;126:430–442.
https://doi.org/10.1016/j.envint.2019.02.042
25. Seyoum MM, Obayomi O, Bernstein N, et al. Occurrence and distribution of antibiotics and corresponding antibiotic resistance genes in different soil types irrigated with treated wastewater. Sci. Total Environ. 2021;782:146835.
https://doi.org/10.1016/j.scitotenv.2021.146835
26. Kuang Y, Guo X, Hu J, et al. Occurrence and risks of antibiotics in an urban river in northeastern Tibetan Plateau. Sci Rep. 2020. 10:20054.
https://doi.org/10.1038/s41598-020-77152-5
27. Jang J, Park J, Hwang CY, et al. Abundance and diversity of antibiotic resistance genes and bacterial communities in the western Pacific and Southern Oceans. Sci. Total Environ. 2022;822:153360.
https://doi.org/10.1016/j.scitotenv.2022.153360
28. Toraskar AD, Manohar CS, Fernandes CL, et al. Seasonal variations in the water quality and antibiotic resistance of microbial pollution indicators in the Mandovi and Zuari estuaries, Goa, India. Environ. Monitor. Assess. 2022. 194:71.
https://doi.org/10.1007/s10661-021-09679-7
29. Naquin A, Shrestha A, Sherpa M, et al. Presence of antibiotic resistance genes in a sewage treatment plant in Thibodaux, Louisiana, USA. Bioresour. Technol. 2015;188:79–83.
https://doi.org/10.1016/j.biortech.2015.01.052
30. Zhang X, Xu Y, He X, et al. Occurrence of antibiotic resistance genes in landfill leachate treatment plant and its effluent-receiving soil and surface water. Environ. Pollut. 2016;218:1255–1261.
https://doi.org/10.1016/j.envpol.2016.08.081
31. Lanyon CW, King JR, Stekel DJ, et al. A Model to Investigate the Impact of Farm Practice on Antimicrobial Resistance in UK Dairy Farms. Bull. Math. Biol. 2021;83:36.
https://doi.org/10.1007/s11538-021-00865-9
32. Antillon GO, Sjöström K, Fall N, et al. Antibiotic Use in Organic and Non-Organic Swedish Dairy Farms: A Comparison of Three Recording Methods. Front. Vet. Sci. 2020;7:568881.
https://doi.org/10.3389/fvets.2020.568881
33. Kokou F, Sasson G, Mizrahi I, et al. Antibiotic effect and microbiome persistence vary along the European seabass gut. Sci. Rep. 2020;10:10003.
https://doi.org/10.1038/s41598-020-66622-5
34. Xu J, Sangthong R, McNeil E, et al. Antibiotic use in chicken farms in northwestern China. Antimicrob. Resist. Infect. Control. 2020;9:10.
https://doi.org/10.1186/s13756-019-0672-6
35. Joseph SM, Battaglia T, Maritz JM, et al. Longitudinal comparison of bacterial diversity and antibiotic resistance genes in New York city sewage. mSystems. 2019. 44e00327–19.
https://doi.org/10.1128%2FmSystems.00327-19
36. Khan FA, Soderquist B, Jass J. Prevalence and Diversity of Antibiotic Resistance Genes in Swedish Aquatic Environments Impacted by Household and Hospital Wastewater. Front. Microbiol. 2019;0:688.
https://doi.org/10.3389/fmicb.2019.00688
37. Tao Y, Yue Y, Wang J. Abundance and diversity of antibiotic resistance genes possibly released to ambient air by experiments in biology laboratories. Sci. Total Environ. 2021;79:49147.
https://doi.org/10.1016/j.scitotenv.2021.149147
38. Kümmerer K. Significance of antibiotics in the environment. J. Antimicrob. Chemother. 2003;52(1)5–7.
https://doi.org/10.1093/jac/dkg293
39. Kolpin DW, Skopec M, Meyer MT, et al. Urban contribution of pharmaceuticals and other organic wastewater contaminants to streams during differing flow conditions. Sci. Total Environ. 2004;28(1–3)119–130.
https://doi.org/10.1016/j.scitotenv.2004.01.015
40. Hari AC, Paruchuri RA, Sabatini DA, et al. Effects of pH and cationic and nonionic surfactants on the adsorption of pharmaceuticals to a natural aquifer material. Environ. Sci. Technol. 2005;39(8)2592–2598.
https://doi.org/10.1021/es048992m
41. Batt AL, Bruce IB, Aga DS. Evaluating the vulnerability of surface waters to antibiotic contamination from varying waste-water treatment plant discharges. Environ. Pollut. 2006;142(2)295–302.
https://doi.org/10.1016/j.envpol.2005.10.010
42. Mclnnes RS, McCallum GE, Lamberte LE, et al. Horizontal transfer of antibiotic resistance genes in the human gut microbiome. Curr. Opin. Microbiol. 2020;53:35–43.
https://doi.org/10.1016/j.mib.2020.02.002
43. Yadav S, Kapley A. Antibiotic resistance: Global health crisis and metagenomics. Biotechno. Rep. 2021;29:e00604.
https://doi.org/10.1016/j.btre.2021.e00604
44. O’Neill J. Review on Antimicrobial Resistance Antimicrobial Resistance: Tackling a crisis for the health and wealth of nations. London: Review on Antimicrobial Resistance; 2014. Available from: https://www.who.int/news/item/29-04-2019-new-report-calls-for-urgent-action-to-avert-antimicrobial-resistance-crisis
Accessed on June 4th, 2022
45. Xu L, Zhang H, Xiong P. Occurrence, fate, and risk assessment of typical tetracycline antibiotics in the aquatic environment: A review. Sci. Total Environ. 2021;753:141975.
https://doi.org/10.1016/j.scitotenv.2020.141975
46. Cheng D, Xie Y, Yu Y, et al. Occurrence and Partitioning of Antibiotics in the Water Column and Bottom Sediments from the Intertidal Zone in the Bohai Bay, China. Wetlands. 2016;36:167–179.
https://doi.org/10.1007/s13157-014-0561-y
47. Hu Y, Yan X, Shen Y, et al. Antibiotics in surface water and sediments from Hanjiang River, Central China: Occurrence, behavior and risk assessment. Ecotoxicol. Environ. Saf. 2018;157:150–158.
https://doi.org/10.1016/j.ecoenv.2018.03.083
48. Taneja N, Sharma M. Antimicrobial resistance in the environment: The Indian scenario. Indian J. Med. Res. 2019;149(2)119–128.
https://doi.org/10.4103%2Fijmr.IJMR_331_18
49. Yan C, Yang Y, Zhou J, et al. Antibiotics in the surface water of the Yangtze Estuary: Occurrence, distribution and risk assessment. Environ. Pollut. 2013;175:22–29.
https://doi.org/10.1016/j.envpol.2012.12.008
50. Shi H, Yang Y, Liu M, et al. Occurrence and distribution of antibiotics in the surface sediments of the Yangtze Estuary and nearby coastal areas. Mar. Pollut. Bull. 2014;83(1)317–323.
https://doi.org/10.1016/j.marpolbul.2014.04.034
51. Zheng S, Qiu X, Chen B, et al. Antibiotics pollution in Jiulong River estuary: Source, distribution and bacterial resistance. Chemosphere. 2011;84(11)1677–1685.
https://doi.org/10.1016/j.chemosphere.2011.04.076
52. Silvester R, Alexander D, Ammanamveetil MHA. Prevalence, antibiotic resistance, virulence and plasmid profiles of Vibrio parahaemolyticus from a tropical estuary and adjoining traditional prawn farm along the southwest coast of India. Ann. Microbiol. 2015;65:2141–2149.
https://doi.org/10.1007/s13213-015-1053-x
53. Na G, Lu Z, Gao H, et al. The effect of environmental factors and migration dynamics on the prevalence of antibiotic-resistant Escherichia coli in estuary environments. Sci. Rep. 2018;8:1663.
https://doi.org/10.1038/s41598-018-20077-x
54. Ohore OE, Zhang S, Guo S, et al. Ciprofloxacin increased abundance of antibiotic resistance genes and shaped microbial community in epiphytic biofilm on Vallisneria spiralis in mesocosmic wetland. Bioresour. Technol. 2021;3(23)124574.
https://doi.org/10.1016/j.biortech.2020.124574
55. Ryan PG, Moore CJ, van Franeker JA, et al. Monitoring the abundance of plastic debris in the marine environment. Philos. Trans. R. Soc. B. 2009;364:1999–2012.
https://doi.org/10.1098/rstb.2008.0207
56. Peeken I, Primpke S, Beyer B, et al. Arctic sea ice is an important temporal sink and means of transport for microplastic. Nat. Commun. 2018;9:505.
https://doi.org/10.1038%2Fs41467-018-03825-5
57. Sheela AM, Manimekalai B, Dhinagaran G. Review on the distribution of microplastics in the oceans and its impacts: Need for modeling-based approach to investigate the transport and risk of microplastic pollution. Environ. Eng. Res. 2022;27(4)210243.
https://doi.org/10.4491/eer.2021.243
58. Kawecki D, Nowack B. Polymer-Specific Modeling of the Environmental Emissions of Seven Commodity Plastics as Macro- and Microplastics. Environ. Sci. Technol. 2019;53(16)9664–9676.
https://doi.org/10.1021/acs.est.9b02900
59. Kumar MV, Sheela AM. Effect of plastic film mulching on the distribution of plastic residues in agricultural fields. Chemosphere. 2021;273:128590.
https://doi.org/10.1016/j.chemosphere.2020.128590
60. van den Berg P, Huerta - Lwanga E, Corradini F, et al. Sewage sludge application as a vehicle for microplastics in eastern Spanish agricultural soils. Environ. Pollut. 2020;261:114198.
https://doi.org/10.1016/j.envpol.2020.114198
61. Franco AA, Arellano JM, Albendin G, et al. Microplastic pollution in wastewater treatment plants in the city of Cádiz: Abundance, removal efficiency and presence in receiving water body. Sci. Total Environ. 2021;776:145795.
https://doi.org/10.1016/j.scitotenv.2021.145795
62. Wan Y, Chen X, Liu Q, et al. Informal landfill contributes to the pollution of microplastics in the surrounding environment. Environ. Pollut. 2022;293:118586.
https://doi.org/10.1016/j.envpol.2021.118586
63. Golwala H, Zhang X, Iskander S, et al. Solid waste: An overlooked source of microplastics to the environment. Sci. Total Environ. 2021;769:144581.
https://doi.org/10.1016/j.scitotenv.2020.144581
64. Lwanga EH, Gertsen H, Gooren H, et al. Incorporation of microplastics from litter into burrows of Lumbricusterrestris. Environ. Pollut (Part A). 2017;220:523–531.
https://doi.org/10.1016/j.envpol.2016.09.096
65. Scheurer M, Bigalke M. Microplastics in Swiss Floodplain Soils. Environ. Sci. Technol. 2018;52(6)3591–3598.
https://doi.org/10.1021/acs.est.7b06003
66. Napper IE, Davies BFR, Clifford H, et al. Reaching New Heights in Plastic Pollution—Preliminary Findings of Microplastics on Mount Everest. One Earth. 2020. 5:621–630.
http://dx.doi.org/10.1016/j.oneear.2020.10.020
67. Fuller S, Gautam AA. Procedure for measuring microplastics using pressurized fluid extraction. Environ. Sci. Technol. 2016;50:5774–5780.
https://doi.org/10.1021/acs.est.6b00816
68. Pabortsava K, Lampitt RS. High concentrations of plastic hidden beneath the surface of the Atlantic Ocean. Nat. Commun. 2020;11:4073.
https://doi.org/10.1038/s41467-020-17932-9
69. Hossain S, Rahman MA, Chowdhury M, Mohonta SK. Plastic pollution in Bangladesh: A review on current statusemphasizing the impacts on environment and public health. Environ. Eng. Res. 2021;26(6)200535.
https://doi.org/10.4491/eer.2020.535
70. Tanentzap AJ, Cottingham S, Fonvielle J, et al. Microplastics and anthropogenic fibre concentrations in lakes reflect surrounding land use. PLoS Biol. 2021;19(9)e3001389.
https://doi.org/10.1371/journal.pbio.3001389
71. Li J, Huang W, Jiang R, et al. Are bacterial communities associated with microplastics influenced by marine habitats? Sci. Total Environ. 2020;733:139400.
https://doi.org/10.1016/j.scitotenv.2020.139400
72. Melvin J, Bury M, Ammendolia J, et al. Critical Gaps in Shoreline Plastics Pollution Research. Front. Mar. Sci. 2021;8:689108.
https://doi.org/10.3389/fmars.2021.689108
73. Wu N, Zhang Y, Zhang X. Occurrence and distribution of microplastics in the surface water and sediment of two typical estuaries in Bohai Bay, China. Environ. Sci. Process. Imp. 2019;21:1143–1152.
https://doi.org/10.1039/C9EM00148D
74. Talley TS, Venuti N, Whelan R. Natural history matters: Plastics in estuarine fish and sediments at the mouth of an urban watershed. PLoS One. 2020;15(3)e0229777.
https://doi.org/10.1371/journal.pone.0229777
75. Lu I, Chao H, Mansor WNW. Levels of Phthalates, Bisphenol-A, Nonylphenol, and Microplastics in Fish in the Estuaries of Northern Taiwan and the Impact on Human Health. Toxics. 2021;9(10)246.
https://doi.org/10.3390/toxics9100246
76. Willis K, Eriksen R, Wilcox C. Microplastic Distribution at Different Sediment Depths in an Urban Estuary. Front. Mar. Sci. 2017;4:419.
https://doi.org/10.3389/fmars.2017.00419
77. Radhakrishnan K, Sivapriya V, Rajkumar , et al. Characterization and distribution of microplastics in estuarine surface sediments, Kayamkulam estuary, southwest coast of India. Mar. Pollut. Bull. 2021;168:112389.
https://doi.org/10.1016/j.marpolbul.2021.112389
78. Li S, Wang Y, Liu L, et al. Temporal and Spatial Distribution of Microplastics in a Coastal Region of the Pearl River Estuary, China. Water. 2021;13(12)1618.
https://doi.org/10.3390/w13121618
79. Lechthaler S, Waldschläger K, Sandhani CG, et al. Baseline Study on Microplastics in Indian Rivers under Different Anthropogenic Influences. Water. 2021;13:1648.
https://doi.org/10.3390/w13121648
80. Sruthy S, Ramasamy EV. Microplastic pollution in Vembanad Lake, Kerala, India: The first report of microplastics in lake and estuarine sediments in India. Environ. Pollut. 2017;222:315–322.
https://doi.org/10.1016/j.envpol.2016.12.038
81. Lima ARA, Costa MF, Barletta M. Distribution patterns of microplastics within the plankton of a tropical estuary. Environ Res. 2014;132:146–155.
https://doi.org/10.1016/j.envres.2014.03.031
82. Zaki MRM, Zaid SHM, Zainuddin AH, et al. Microplastic pollution in tropical estuary gastropods: Abundance, distribution and potential sources of Klang River estuary, Malaysia. Mar. Pollut. Bull. 2021;162:111866.
https://doi.org/10.1016/j.marpolbul.2020.111866
83. Zhao S, Zhu L, Li D. Microplastic in three urban estuaries, China. Environ. Pollut. 2015;206:597–604.
https://doi.org/10.1016/j.envpol.2015.08.027
84. Peng G, Zhu B, Yang D, et al. Microplastics in sediments of the Changjiang Estuary, China. Environ. Pollut. 2017;225:283–290.
https://doi.org/10.1016/j.envpol.2016.12.064
85. Gray AD, Wertz H, Leads RR, et al. Microplastic in two South Carolina Estuaries: Occurrence, distribution, and composition. Mar. Pollut. Bull. 2018;128:223–233.
https://doi.org/10.1016/j.marpolbul.2018.01.030
86. Ta AT, Babel S. Microplastics pollution with heavy metals in the aquaculture zone of the Chao Phraya River Estuary, Thailand. Mar. Pollut. Bull(Part A). 2020;161:111747.
https://doi.org/10.1016/j.marpolbul.2020.111747
87. Pan Z, Liu Q, Jiang R, et al. Microplastic pollution and ecological risk assessment in an estuarine environment: The Dongshan Bay of China. Chemosphere. 2021;262:127876.
https://doi.org/10.1016/j.chemosphere.2020.127876
88. Diaz-Jaramillo M, Islas MS, Gonzalez M. Spatial distribution patterns and identification of microplastics on intertidal sediments from urban and semi-natural SW Atlantic estuaries. Environ. Pollut. 2021;273:116398.
https://doi.org/10.1016/j.envpol.2020.116398
89. Gupta P, Saha M, Rathore C. Spatial and seasonal variation of microplastics and possible sources in the estuarine system from central west coast of India. Environ. Pollut. 2021;288:117665.
https://doi.org/10.1016/j.envpol.2021.117665
90. Hitchcock JN, Mitrovic SM. Microplastic pollution in estuaries across a gradient of human impact. Environ. Pollut. 2019;247:457–466.
https://doi.org/10.1016/j.envpol.2019.01.069
91. Kerubo J, Muthumbi A, Onyari JM, et al. Microplastics pollution in the sediments of creeks and estuaries of Kenya, western Indian Ocean. Afr. J. Mar. Sci. 2021;43(3)337–352.
https://doi.org/10.2989/1814232X.2021.1966505
92. Haddout S, Gimiliani GT, Priya KL, et al. Microplastics in Surface Waters and Sediments in the Sebou Estuary and Atlantic Coast, Morocco. Anal Lett. 2021;1–13.
https://doi.org/10.1080/00032719.2021.1924767
93. Neeji N, Rafeei M, Azizi H, et al. Data on the microplastics contamination in water and sediments along the Haraz River estuary, Iran. Data Br. 2020;32:106155.
https://doi.org/10.1016%2Fj.dib.2020.106155
94. Gallagher A, Rees A, Rowe R, et al. Microplastics in the Solent estuarine complex, UK: An initial assessment. Mar. Pollut. Bull. 2016;102(2)243–249.
https://doi.org/10.1016/j.marpolbul.2015.04.002
95. Naidoo T, Glassom D, Smit AJ. Plastic pollution in five urban estuaries of KwaZulu-Natal, South Africa. Mar. Pollut. Bull. 2015;101(1)473–480.
https://doi.org/10.1016/j.marpolbul.2015.09.044
96. Suteja Y, Atmadipoera A, Riani E, et al. Spatial and temporal distribution of microplastic in surface water of tropical estuary: Case study in Benoa Bay, Bali, Indonesia. Mar. Pollut. Bull. 2021;16:111979.
https://doi.org/10.1016/j.marpolbul.2021.111979
97. Forero-Lopez AD, Rimondino GN, Truchet DM, et al. Occurrence, distribution, and characterization of suspended microplastics in a highly impacted estuarine wetland in Argentina. Sci. Total Environ. 2021;785:147141.
https://doi.org/10.1016/j.scitotenv.2021.147141
98. Villagran DM, Truchet DM, Buzzi NS, et al. A baseline study of microplastics in the burrowing crab (Neohelice granulata) from a temperate southwestern Atlantic estuary. Mar. Pollut. Bull. 2020;150:110686.
https://doi.org/10.1016/j.marpolbul.2019.110686
99. Vedolin MC, Teophilo CYS, Turra A, Figueira RCL. Spatial variability in the concentrations of metals in beached microplastics. Mar. Pollut. Bull. 2018;129(2)487–489.
https://doi.org/10.1016/j.marpolbul.2017.10.019
100. Stead JL, Cundy AB, Hudson MD, et al. Identification of tidal trapping of microplastics in a temperate salt marsh system using sea surface microlayer sampling. Sci. Rep. 2020;10:14147.
https://doi.org/10.1038/s41598-020-70306-5
101. Yao W, Di D, Wang Z, et al. Micro and macroplastic accumulation in a newly formed Spartinaalterniflora colonized estuarine saltmarsh in southeast China. Mar. Pollut. Bull. 2019;149:110636.
https://doi.org/10.1016/j.marpolbul.2019.110636
102. Liu P, Zhan X, Wu X, et al. Effect of weathering on environmental behavior of microplastics: Properties, sorption and potential risks. Chemosphere. 2020;242:125193.
https://doi.org/10.1016/j.chemosphere.2019.125193
103. Bailey K, Sipps MK, Saba GK, et al. Quantification and composition of microplastics in the Raritan Hudson Estuary: Comparison to pathways of entry and implications for fate. Chemosphere. 2021;272:129886H.
https://doi.org/10.1016/j.chemosphere.2021.129886
104. Pickett JE. Handbook of Environmental Degradation of Materials. 3rd edOxford: AcademicPress; 2018. p. 163–184.
105. Andrady AL. The plastic in microplastics: A review. Mar. Pollut. Bull. 2017;119(1)12–22.
https://doi.org/10.1016/j.marpolbul.2017.01.082
106. Arp HPH, Kuhnel D, Rummel C, et al. Weathering Plastics as a Planetary Boundary Threat: Exposure, Fate, and Hazards. Environ. Sci. Technol. 2021;55(11)7246–7255.
https://doi.org/10.1021/acs.est.1c01512
107. Chevin LM, Hoffmann AA. Evolution of phenotypic plasticity in extreme environments. Philos. Trans. R. Soc. B. 2017;372:20160138.
https://doi.org/10.1098/rstb.2016.0138
108. Hurtado-Bautista E, Pérez Sánchez LF, Islas-Robles A, et al. Phenotypic plasticity and evolution of thermal tolerance in bacteria from temperate and hot spring environments. PeerJ. 2021;9:e11734.
https://doi.org/10.7717/peerj.11734
109. Hansen L, Ren D, Burmølle M, et al. Distinct gene expression profile of Xanthomonasretroflexus engaged in synergistic multispecies biofilm formation. ISME J. 2017;11:300–303.
https://doi.org/10.1038%2Fismej.2016.107
110. Beech IB. Corrosion of technical materials in the presence of biofilms—current understanding and state-of-the art methods of study. Int. Biodeterior. Biodegradation. 2004;53(3)177–183.
http://dx.doi.org/10.1016%2FS0964-8305(03)00092-1
111. Flemming HC, Wingender J, Szewzyk U, et al. Biofilms: an emergent form of bacterial life. Nat. Rev. Microbiol. 2016;14:563–575.
https://doi.org/10.1038/nrmicro.2016.94
112. Jaques M, Aragon V, Tremblay YDN. Biofilm formation in bacterial pathogens of veterinary importance. Anim. Health Res Rev. 2010;11(2)97–121.
https://doi.org/10.1017/s1466252310000149
113. Costerton JW, Stewart PE, Greenberg EP. Bacterial Biofilms: A Common Cause of Persistent Infections. Science. 1999;284(5418)1318–1322.
https://doi.org/10.1126/science.284.5418.1318
114. Marques VF, da Motta CC, Soares BS, et al. Biofilm production and beta-lactamic resistance in Brazilian Staphylococcus aureus isolates from bovine mastitis. Braz. J. Microbiol. 2007;48(1)118–124.
https://doi.org/10.1016/j.bjm.2016.10.001
115. Zou D, Liu H, Li M. Community, Distribution, and Ecological Roles of Estuarine Archaea. Front. Microbiol. 2020;11:2060.
https://doi.org/10.3389/fmicb.2020.02060
116. Xu YB, Hou MY, Li YF, et al. Distribution of tetracycline resistance genes and AmpC β-lactamase genes in representative non-urban sewage plants and correlations with treatment processes and heavy metals. Chemosphere. 2017;170:274–281.
https://doi.org/10.1016/j.chemosphere.2016.12.02
117. Zhang H, Wang J, Zhou B, et al. Enhanced adsorption of oxytetracycline to weathered microplastic polystyrene: Kinetics, isotherms and influencing factors. Environ. Pollut (Part B). 2018;243:1550–1557.
https://doi.org/10.1016/j.envpol.2018.09.122
118. Chihomvu P, Stegmann P, Pillay M. Characterization and Structure Prediction of Partial Length Protein Sequences of pcoA; pcoR and chrB Genes from Heavy Metal Resistant Bacteria from the Klip River; South Africa. Int. J. Mol. Sci. 2015;16:7352–7374.
https://doi.org/10.3390/ijms16047352
119. Komijani M, Shamabadi NS, Shahin K, et al. Heavy metal pollution promotes antibiotic resistance potential in the aquatic environment. Environ Pollut. 2021;274:116569.
https://doi.org/10.1016/j.envpol.2021.116569
120. Suzuki S, Kimura M, Agusa T, et al. Vanadium accelerates horizontal transfer of tet(M) gene from marine Photobacterium to Escherichia coli
. FEMS Microbiol.Lett. 2012;336:52–56.
https://doi.org/10.1111/j.1574-6968.2012.02653.x
121. Yu F, Yang C, Huang G, et al. Interfacial interaction between diverse microplastics and tetracycline by adsorption in an aqueous solution. Sci.Total Environ. 2020;721:137729.
https://doi.org/10.1016/j.scitotenv.2020.137729
122. Zettler ER, Mincer TJ, Amaral-Zettler LA. Life in the “Plastisphere”: Microbial Communities on Plastic Marine Debris. Environ. Sci. Technol. 2013;47(13)7137–7146.
https://doi.org/10.1021/es401288x
123. Voulgari E, Zarkotou O, Ranellou K, et al. Outbreak of OXA-48 carbapenemase-producing Klebsiellapneumoniae in Greece involving an ST11 clone. J. Antimicrob. Chemother. 2013;68(1)84–88.
https://doi.org/10.1093/jac/dks356
124. Anderi JN, Franklin MJ, Stewart PS. Role of antibiotic penetration limitation in Klebsiellapneumoniae Biofilm Resistance to Ampicillin and Ciprofloxacin. Antimicrob. Agents Chemother. 2020;44(7)1818–1824.
https://doi.org/10.1128%2Faac.44.7.1818-1824.2000
125. Roose-Amsaleg C, Laverman AM. Do antibiotics have environmental side-effects? Impact of synthetic antibiotics on bio-geochemical processes. Environ. Sci. Pollut. Res. 2016;23:4000–4012.
https://doi.org/10.1007/s11356-015-4943-3
126. Sukumaran DP, Durairaj S, Abdulla MH, et al. Antibiotic Resistance of Escherichia coli Serotypes from Cochin Estuary. Interdiscip Perspect Infect Dis. 2012;124879:1–7.
https://doi.org/10.1155/2012/124879
127. Liang X, Guan F, Chen B, et al. Spatial and seasonal variations of antibiotic resistance genes and antibiotics in the surface waters of Poyang Lake in China. Ecotoxicol. Environ Saf. 2020;196:110543.
https://doi.org/10.1016/j.ecoenv.2020.110543
128. Lin L, Yuan K, Liang X, et al. Occurrences and distribution of sulfonamide and tetracycline resistance genes in the Yangtze River Estuary and nearby coastal area. Mar. Pollut. Bull. 2015;100(1)304–310.
https://doi.org/10.1016/j.marpolbul.2015.08.036
129. Laverty AL, Primpke S, Lorenz C, et al. Bacterial biofilms colonizing plastics in estuarine waters, with an emphasis on Vibrio spp. and their antibacterial resistance. PLoS One. 2020;15(8)e0237704.
https://doi.org/10.1371/journal.pone.0237704
130. Zhao Z, Zhang K, Wu N, et al. Estuarine sediments are key hotspots of intracellular and extracellular antibiotic resistance genes: A high-throughput analysis in Haihe Estuary in China. Environ. Int. 2020;135:105385.
https://doi.org/10.1016/j.envint.2019.105385
131. Guo X, Sun X, Chen Y, et al. Antibiotic resistance genes in biofilms on plastic wastes in an estuarine environment. Sci. Total Environ. 2020;745:140916.
https://doi.org/10.1016/j.scitotenv.2020.140916
132. Liang X, Guan F, Chen B, et al. Spatial and seasonal variations of antibiotic resistance genes and antibiotics in the surface waters of Poyang Lake in China. Ecotoxicol. EnvironSaf. 2020;196:110543.
https://doi.org/10.1016/j.ecoenv.2020.110543
133. Lin L, Yuan K, Liang X, et al. Occurrences and distribution of sulfonamide and tetracycline resistance genes in the Yangtze River Estuary and nearby coastal area. Mar. Pollut. Bull. 2015;100(1)304–310.
https://doi.org/10.1016/j.marpolbul.2015.08.036
134. Lu X, Lu P. Seasonal variations in antibiotic resistance genes in estuarine sediments and the driving mechanisms. J Hazard Mater. 2020;383:121164.
http://dx.doi.org/10.1016/j.jhazmat.2019.121164
135. Chen B, Liang X, Huang X, et al. Differentiating anthropogenic impacts on ARGs in the Pearl River Estuary by using suitable gene indicators. Water Res. 2013;47(8)2811–2820.
https://doi.org/10.1016/j.watres.2013.02.042
136. Divya SP, Hatha AAM. Screening of tropical estuarine water in south-west coast of India reveals emergence of ARGs-harboring hypervirulent Escherichia coli of global significance. Int. J. Hyg. Environ. Health. 2019;222(2)235–248.
https://doi.org/10.1016/j.ijheh.2018.11.002
137. Bhattarchyya A, Haldar A, Bhattarchyya M, et al. Anthropogenic influence shapes the distribution of antibiotic resistant bacteria (ARB) in the sediment of Sundarban estuary in India. Sci. Total Environ. 2019;647:1626–1639.
https://doi.org/10.1016/j.scitotenv.2018.08.038
138. Bhandari S, Adhikari S, Karki D, et al. Antibiotic Resistance, Biofilm Formation and Detection of mexA/mexB Efflux-Pump Genes Among Clinical Isolates of Pseudomonas aeruginosa in a Tertiary Care Hospital, Nepal. Front. Trop. Dis. 2022;2:810863.
https://doi.org/10.3389/fitd.2021.810863
139. Christaki E, Marcou M, Tofarides A. Antimicrobial Resistance in Bacteria: Mechanisms, Evolution, and Persistence. J. Mol Evol. 2020;88(1)26–40.
https://doi.org/10.1007/s00239-019-09914-3
140. Jiang H, Cheng H, Liang Y, et al. Diverse Mobile Genetic Elements and Conjugal Transferability of Sulfonamide Resistance Genes (sul1, sul2, and sul3) in Escherichia coli Isolates FromPenaeusvannamei and Pork From Large Markets in Zhejiang, China. Front. Microbiol. 2019;10:1787.
https://doi.org/10.3389/fmicb.2019.01787
141. Sucato A, Vecchioni L, Savoca D, et al. A Comparative Analysis of Aquatic and Polyethylene-Associated Antibiotic-Resistant Microbiota in the Mediterranean Sea. Biology. 2021;10(3)200.
https://doi.org/10.3390/biology10030200
142. Singh S, Datta S, Narayanan KB, et al. Bacterial exo-polysaccharides in biofilms: role in antimicrobial resistance and treatments. J. Genet. Eng. Biotechnol. 2021;9:140.
https://doi.org/10.1186/s43141-021-00242-y
143. Lin Z, Zhou Z, Zhu L, et al. Behavior of antibiotic resistance genes in a wastewater treatment plant with different upgrading processes. Sci. Total Environ. 2021;771:44814.
https://doi.org/10.1016/j.scitotenv.2020.144814
144. Chikwendu CI, Ibe SN, Okpokwasili GC. Detection of blaSHV and blaTEM beta-lactamase genes in multi-resistant Pseudomonas isolates from environmental sources. Afr. J. Microbiol. Res. 2011;5(15)2067–2074.
https://doi.org/10.5897/AJMR11.149
145. Li H, Song R, Wang Y, et al. Inhibited conjugative transfer of antibiotic resistance genes in antibiotic resistant bacteria by surface plasma. Water Res. 2021;204:117630.
https://doi.org/10.1016/j.watres.2021.117630
146. Chen B, Hao L, Guo X, et al. Prevalence of antibiotic resistance genes of wastewater and surface water in livestock farms of Jiangsu Province, China. Environ. Sci. Pollut. Res. 2015;22:13950–13959.
https://doi.org/10.1007/s11356-015-4636-y
147. Hedayatianfard K, Akhlaghi M, Sharifiyazdi . Detection of tetracycline resistance genes in bacteria isolated from fish farms using polymerase chain reaction. Vet. Res. Forum. 2014;5(4)269–275.
148. Kallau NHG, Wibawan WT, Lukman DW, et al. Detection of multi-drug resistant (MDR) Escherichia coli and tet gene prevalence at a pig farm in Kupang, Indonesia. J. Adv. Vet. Anim. Res. 2018;5(4)388–396.
https://doi.org/10.5455%2Fjavar.2018.e289
149. Ahmed MAEE, Shen LZC, Yang Y, et al. Colistin and its role in the Era of antibiotic resistance: an extended review (2000–2019). Emerg. Microbes & Infect. 2020;9(1)868–885.
https://doi.org/10.1080/22221751.2020.1754133
150. Rezazadeh M, Baghchesaraei H, Peymani A. Plasmid-Mediated Quinolone-Resistance (qnr) Genes in Clinical Isolates of Escherichia coli Collected from Several Hospitals of Qazvin and Zanjan Provinces, Iran. Osong Public Health Res. Perspect. 2016;7(5)307–312.
https://doi.org/10.1016%2Fj.phrp.2016.08.003
151. Abu AB, Mirabeau TY. Prevalence of qnr Genes among Multidrug Resistance Staphylococcus aureus from Clinical Isolates. J. Adv. Med. Med. Res. 2019;30(10)1–10.
https://doi.org/10.9734/jammr/2019/v30i1030245
152. Alder J. Putting the Coast in the Sea around Us Project. The Sea around Us Newsletter. 152003;1–2.
153. Marchand S, Block JD, Jonghe VD, et al. Biofilm Formation in Milk Production and Processing Environments; Influence on Milk Quality and Safety. Compr Rev Food Sci. 2012;1(2)133–147.
https://doi.org/10.1111/j.1541-4337.2011.00183.x
154. Roy PK, Ji-Won-Ha J, Mizan MFR, et al. Effects of environmental conditions (temperature, pH, and glucose) on biofilm formation of Salmonella enterica serotype Kentucky and virulence gene expression. Poult Sci J. 2021;100(7)101209.
https://doi.org/10.1016%2Fj.psj.2021.101209
155. Lopez D, Vlamakis H, Kolter R. Biofilms. Cold Spring Harb. Perspect. Biol. 2010;2:a000398.
https://doi.org/10.1101/cshperspect.a000398
156. Doghri I, Rodrigues S, Bazire A, et al. Marine bacteria from the French Atlantic coast displaying high forming-biofilm abilities and different biofilm 3D architectures. BMC Microbiol. 2015;15:231.
https://doi.org/10.1186/s12866-015-0568-4
157. Jones PR, Cottrell MT, Kirchman DL, et al. Bacterial Community Structure of Biofilms on Artificial Surfaces in an Estuary. Microb. Ecol. 2006;53:153–162.
https://doi.org/10.1007/s00248-006-9154-5
158. Guo X, Niu Z, Lu D, et al. Bacterial community structure in the intertidal biofilm along the Yangtze Estuary, China. Mar. Pollut. Bull. 2017;124(1)314–320.
https://doi.org/10.1016/j.marpolbul.2017.07.051
159. Jiang PL, Zhao SY, Zhu LX, et al. Microplastic-associated bacterial assemblages in the intertidal zone o the Yangtze Estuary. Sci. Total Environ. 2018;624:48–54.
https://doi.org/10.1016/j.scitotenv.2017.12.105
160. Mai YZ, Peng SY, Lai ZN. Structural and functional diversity of biofilm bacterial communities along the Pearl River Estuary, South China. Reg. Stud. Mar. Sci. 2020;33:100926.
http://doi.org/10.1016/j.rsma.2019.100926
161. Wu N, Zhang Y, Zhao Z, et al. Colonization characteristics of bacterial communities on microplastics compared with ambient environments (water and sediment) in Haihe Estuary. Sci. Total Environ. 2020;708:134876.
https://doi.org/10.1016/j.scitotenv.2019.134876
162. Xu X, Wang S, Gao F, et al. Marine microplastic-associated bacterial community succession in response to geography, exposure time, and plastic type in China’s coastal seawaters. Mar. Pollut. Bull. 2019;145:278–286.
163. Tu C, Chen T, Zhou Q, et al. Biofilm formation and its influences on the properties of microplastics as affected by exposure time and depth in the seawater. Sci. Total Environ. 2020;734:139237.
https://doi.org/10.1016/j.scitotenv.2020.139237
164. Zhang B, Yang X, Liu L, et al. Spatial and seasonal variations in biofilm formation on microplastics in coastal waters. Sci. Total Environ. 2021;770:145303.
https://doi.org/10.1016/j.scitotenv.2021.145303
165. Lear L, Padfield D, Dowsett T, et al. Bacterial colonisation dynamics of household plastics in a coastal environment. Sci. Total Environ. 2022;838(Part4)156199.
https://doi.org/10.1016/j.scitotenv.2022.156199
166. Issac MN, Kandasubramanian B. Effect of microplastics in water and aquatic systems. Environ. Sci. Pollut. Res. 2021;28:19544–19562.
https://doi.org/10.1007/s11356-021-13184-2
167. Wang T, Hu M, Song L, et al. Coastal zone use influences the spatial distribution of microplastics in Hangzhou Bay, China. Environ. Pollut(Part 2). 2020;266:115137.
https://doi.org/10.1016/j.envpol.2020.115137
168. Else TN, Pantle CR, Amy P. Boundaries for biofilm formation: Humidity and temperature. Appl. Environ. Microbiol. 2020;69(8)5006–5010.
https://doi.org/10.1128%2FAEM.69.8.5006-5010.2003
169. Li R, Yu L, Chai M, et al. The distribution, characteristics and ecological risks of microplastics in the mangroves of Southern China. Sci. Total Environ. 2020;708:135025.
https://doi.org/10.1016/j.scitotenv.2019.135025
170. Rochex A, Lebeault JM. Effects of nutrients on biofilm formation and detachment of a Pseudomonas putida strain isolated from a paper machine. Water Res. 2007;41(13)2885–2892.
https://doi.org/10.1016/j.watres.2007.03.041
171. Kim KY, Frank JF. Effect of Nutrients on Biofilm Formation by Listeria monocytogenes on Stainless Steel. J. Food Prot. 1995;58(1)24–28.
https://doi.org/10.4315/0362-028x-58.1.24
172. Dang H, Lovell CR. Microbial surface colonization and biofilm development in marine environments. Microbiol. Mol. Biol. Rev. 2016;8(1)91–138.
https://doi.org/10.1128/mmbr.00037-15
173. Skovhus TL, Eckert RB, Rodrigues E. Management and control of microbiologically influenced corrosion (MIC) in the oil and gas industry—Overview and a North Sea case study. J Biotechnol. 2017;256:31–45.
https://doi.org/10.1016/j.jbiotec.2017.07.003
174. Balcazar J, Subirats J, Borrego CM. The role of biofilms as environmental reservoirs of antibiotic resistance. Front. Microbiol. 2015;6:1216.
https://doi.org/10.3389/fmicb.2015.01216
175. Zhang B, Yang X, Liu L, et al. Spatial and seasonal variations in biofilm formation on microplastics in coastal waters. Sci. Total Environ. 2021;770:145303.
https://doi.org/10.1016/j.scitotenv.2021.145303
176. Miao L, Wang P, Hou J, et al. Distinct community structure and microbial functions of biofilms colonizing microplastics. Sci Total Environ(Part 2). 2019;650:2395–2402.
https://doi.org/10.1016/j.scitotenv.2018.09.378
177. Debroas D, Mone A, Halle AT. Plastics in the North Atlantic garbage patch: A boat-microbe for hitchhikers and plastic degraders. Sci Total Environ. 2017;599–600:1222–1232.
https://doi.org/10.1016/j.scitotenv.2017.05.059
178. Kesy K, Oberbeckmann S, Kreikemeyer B, et al. Spatial Environmental Heterogeneity Determines Young Biofilm Assemblages on Microplastics in Baltic Sea Mesocosms. Front. Microbiol. 2019;10:1665.
https://doi.org/10.3389/fmicb.2019.01665
179. Saigar-Chaparro SJ, Lepkova K, Pojtnabuntoeng T, et al. Nutrient level determines biofilm characteristics and subsequent impact on microbial corrosion and biocide effectiveness. Appl. Environ. Microbiol. 2020;86:e02885.
https://doi.org/10.1128%2FAEM.02885-19
180. Anwar H, Strap JL, Costerton JW. Establishment of Aging Biofilms: Possible Mechanism of Bacterial Resistance to Antimicrobial Therapy. Antimicrob. Agents Chemother. 1992;36(7)1347–1351.
https://doi.org/10.1128/aac.36.7.1347
181. Qiang L, Cheng J, Mirzoyan S, et al. Characterization of Microplastic-Associated Biofilm Development along a Freshwater-Estuarine Gradient. Environ. Sci. Technol. 2021;55(24)16402–16412.
https://doi.org/10.1021/acs.est.1c04108
182. Jacquin J, Cheng J, Odobel C, et al. Microbial Ecotoxicology of Marine Plastic Debris: A Review on Colonization and Biodegradation by the “Plastisphere”. Front. Microbiol. 2019;10:865.
https://doi.org/10.3389/fmicb.2019.00865
183. Vaksmaa A, Knittel K, Asbun AAA, et al. Microbial Communities on Plastic Polymers in the Mediterranean Sea. Front Microbiol. 2021;12:673553.
https://doi.org/10.3389%2Ffmicb.2021.673553
184. Santos SM, Carbajo JM, Villar JC. The effect of carbon and nitrogen sources on bacterial cellulose production and properties from Gluconacetobactersucrofermentans CECT7291 focused on its use in degraded paper restoration. Bioresources. 2013;8(3)3630–3645.
https://doi.org/10.15376/BIORES.8.3.3630-3645
185. Mehmood CT, Quazi I, Hashmi I, et al. Biodegradation of low density polyethylene (LDPE) modified with dye sensitized titania and starch blend using Stenotrophomonaspavanii
. Int. Biodeterior. Biodegradation. 2020;113:276–286.
http://dx.doi.org/10.1016%2Fj.ibiod.2016.01.025
186. Tu C, Chen T, Zhou Q, et al. Biofilm formation and its influences on the properties of microplastics as affected by exposure time and depth in the seawater. Sci. Total Environ. 2020;734:139237.
https://doi.org/10.1016/j.scitotenv.2020.139237
187. Ogonowski M, Motiei A, Ininbergs K, et al. Evidence for selective bacterial community structuring on microplastics. Environ Microbiol. 2018;20(8)2796–2808.
https://doi.org/10.1111/1462-2920.14120
188. Ramperger AFRM, Stellwang AC, Caspari A, et al. Structural Diversity in Early-Stage Biofilm Formation on Microplastics Depends on Environmental Medium and Polymer Properties. Water. 2020;12(11)3216.
https://doi.org/10.3390/w12113216
189. Miao L, Wang P, Hou J, et al. Distinct community structure and microbial functions of biofilms colonizing microplastics. Sci Total Environ(Part 2). 2019;650:2395–2402.
https://doi.org/10.1016/j.scitotenv.2018.09.378
190. Li Y, Zhang Y, Su F, et al. Adsorption behaviour of microplastics on the heavy metal Cr(VI) before and after ageing. Chemosphere. 2022;302:134865.
https://doi.org/10.1016/j.chemosphere.2022.134865
191. Jeong S, Jang S, Kim SS, et al. Size-dependent seizurogenic effect of polystyrene microplastics in zebrafish embryos. J. Hazard. Mater. 2022;439:129616.
https://doi.org/10.1016/j.jhazmat.2022.129616
192. Nishida H, Tokiwa Y. Distribution of polu(β-hydroxybuty-rate) and poly(ɛ-caprolactone) aerobic degrading microorganisms in different environments. J. Environ. Polym. Degr. 1993;1:227–233.
https://doi.org/10.1007/BF01458031
193. Yoshida S, Hiraga K, Takehana T, et al. A bacterium that degrades and assimilates poly (ethylene terephthalate). Science. 2016;51(6278)1196–1199.
https://doi.org/10.1126/science.aad6359
194. Delacuvellerie A, Cyriaque V, Gobert S, et al. The plastisphere in marine ecosystem hosts potential specific microbial degraders including Alcanivorax borkumensis as a key player for the low-density polyethylene degradation. J. Hazard. Mater. 2019;380:120899.
https://doi.org/10.1016/j.jhazmat.2019.120899
195. Paco A, Duarte K, da Costa JP, et al. Biodegradation of polyethylene microplastics by the marine fungus Zelerion maritimum
. Sci. Total Environ. 2017;586:10–15.
https://doi.org/10.1016/j.scitotenv.2017.02.017
196. Sangeetha Devi R, Kanna VR, Nivas D, et al. Biodegradation of HDPE by Aspergillus spp. from marine ecosystem of Gulf of Mannar, India. Mar. Pollut. Bullet. 2015;96(1–2)32–40.
https://doi.org/10.1016/j.marpolbul.2015.05.050
197. Wang J, Guo X, Xue J. Biofilm-Developed MicroplasticsAs Vectors of Pollutants in Aquatic Environments. Environ. Sci. Technol. 2021;55(1)12780–12790.
https://doi.org/10.1021/acs.est.1c04466
198. Gong M, Yang G, Zhuang L, et al. Microbial biofilm formation and community structure on low-density polyethylene microparticles in lake water microcosms. Environ Pollut. 2019;252(Part2)94–102.
https://doi.org/10.1016/j.envpol.2019.05.090A
199. Abe K, Nomura N, Suzuki S. Biofilms: hot spots of horizontal gene transfer (HGT) in aquatic environments, with a focus on a new HGT mechanism. FEMS Microbiol. Ecol. 2020. 965fiaa031.
https://doi.org/10.1093/femsec/fiaa031
200. Sobecky PA, Hazen TH. Horizontal Gene Transfer and mobile genetic elements in marine systems. Methods Mol. Biol. 2009;532:435–53.
https://doi.org/10.1007/978-1-60327-853-9_25
201. Chiura HX, Kogure K, Hagemann S, et al. Evidence for particle- induced horizontal gene transfer and serial transduction between bacteria. FEMS Microbiol. Ecol. 2011;76(3)576–591.
https://doi.org/10.1111/j.1574-6941.2011.01077.x
202. Zhao Z, Li J, Xue B, et al. Horizontal transfer of the multidrug resistance plasmid RP4 inhibits ammonia nitrogen removal dominated by ammonia-oxidizing bacteria. Water Res. 2022;118434.
https://doi.org/10.1016/j.watres.2022.118434
203. Alawi M, Torrijos TV, Walsh F. Plasmid-mediated antimicrobial resistance in drinking water. Environ. Adv. 2022;8:100191.
https://doi.org/10.1016/j.envadv.2022.100191
204. Zheng D, Yin G, Liu M, et al. A systematic review of antibiotics and antibiotic resistance genes in estuarine and coastal environments. Sci.Total Environ. 2021;777:46009.
https://doi.org/10.1016/j.scitotenv.2021.146009
205. Modi SR, Collins JJ, Relman DA. Antibiotics and the gut microbiota. J. Clin. Investig. 2014;124(10)4212–4218.
https://doi.org/10.1172/jci72333
206. Ma H, Bryers JD. Non-invasive determination of conjugative transfer of plasmids bearing antibiotic-resistance genes in biofilm-bound bacteria: effects of substrate loading and antibiotic selection. Appl. Microbiol. Biotechnol. 2012;97:317–328.
https://doi.org/10.1007/s00253-012-4179-9
207. Molin S, Tolker Nielson T. Gene transfer occurs with enhanced efficiency in biofilms and induces enhanced stabilisation of the biofilm structure. Curr. Opin. Biotechnol. 2003;14(3)255–261.
https://doi.org/10.1016/s0958-1669(03)00036-3
208. Qiu Y, Zhang J, Li B, et al. A novel microfluidic system enables visualization and analysis of antibiotic resistance gene transfer to activated sludge bacteria in biofilm. Sci. Total Environ. 2018;642:582–590.
https://doi.org/10.1016/j.scitotenv.2018.06.012
209. Jiang SC, Paul JH. Gene Transfer by Transduction in the Marine Environment. Appl. Environ. Microbiol. 1998;64(8)2780–2787.
https://doi.org/10.1128/aem.64.8.2780-2787.1998
210. Yu N, Guo B, Liu Y. Shaping biofilm microbiomes by changing GAC location during wastewater anaerobic digestion. Sci. Total Environ. 2021;780:146488.
https://doi.org/10.1016/j.scitotenv.2021.146488
211. Solheim HT, Sekse C, Urdahl AM, et al. Biofilm as an Environment for Dissemination of stx Genes by Transduction. Appl. Environ. Microbiol. 2013;79(3)896–900.
https://doi.org/10.1128%2FAEM.03512-12
212. Syranidou E, Kalogerakis N. Interactions of microplastics, antibiotics and antibiotic resistant genes within WWTPs. Sci. Total Environ. 2022;804:150141.
https://doi.org/10.1016/j.scitotenv.2021.150141
213. Romero JL, Grande Burgos MJ, Pérez-Pulido R, et al. Resistance to Antibiotics, Biocides, Preservatives and Metals in Bacteria Isolated from Seafoods:Co-Selection of Strains Resistant or tolerant to Different Classes of Compounds. Front. Microbiol. 2017;8:1650.
https://doi.org/10.3389/fmicb.2017.01650
214. He L, He L, Gao F, et al. Antibiotics, antibiotic resistance genes and microbial community in grouper mariculture. Sci. Total Environ. 2022;808:152042.
https://doi.org/10.1016/j.scitotenv.2021.152042
215. Ndi OL, Barton MD. Incidence of class 1 integron and other antibiotic resistance determinants in Aeromonas spp. from rainbow trout farms in Australia. J. Fish Dis. 2011;34(8)589–599.
https://doi.org/10.1111/j.1365-2761.2011.01272.x
216. Helsens N, Calvez S, Prevost H, et al. Antibiotic Resistance Genes and Bacterial Communities of Farmed Rainbow Trout Fillets (Oncorhynchus mykiss). Front. Microbiol. 2020;11:590902.
https://doi.org/10.3389/fmicb.2020.590902
217. Miller RA, Harbottle H. Antimicrobial Drug Resistance in Fish Pathogens. Microbiol Spectr. 2018;6(1)
https://doi.org/10.1128/microbiolspec.arba-0017-2017
218. Zdanowicz M, Mudryk ZJ, Perliński P. Abundance and antibiotic resistance of Aeromonas isolated from the water of three carp ponds. Vet. Res. Commun. 2020;44:9–18.
https://doi.org/10.1007/s11259-020-09768-x
219. Yuan L, Wang L, Li Z, et al. Antibiotic resistance and microbiota in the gut of Chinese four major freshwater carp from retail markets. Environ. Pollut(Part 2). 2019;255:113327.
https://doi.org/10.1016/j.envpol.2019.113327
220. Stalin N, Srinivasan P. Molecular characterization of antibiotic resistant Vibrio harveyi isolated from shrimp aquaculture environment in the south east coast of India. Microb. Pathog. 2016;97:110–118.
https://doi.org/10.1016/j.micpath.2016.05.021
Fig. 1Studies conducted to examine microplastic based biofilm formation in estuaries across the world (The red dots indicate the estuaries). 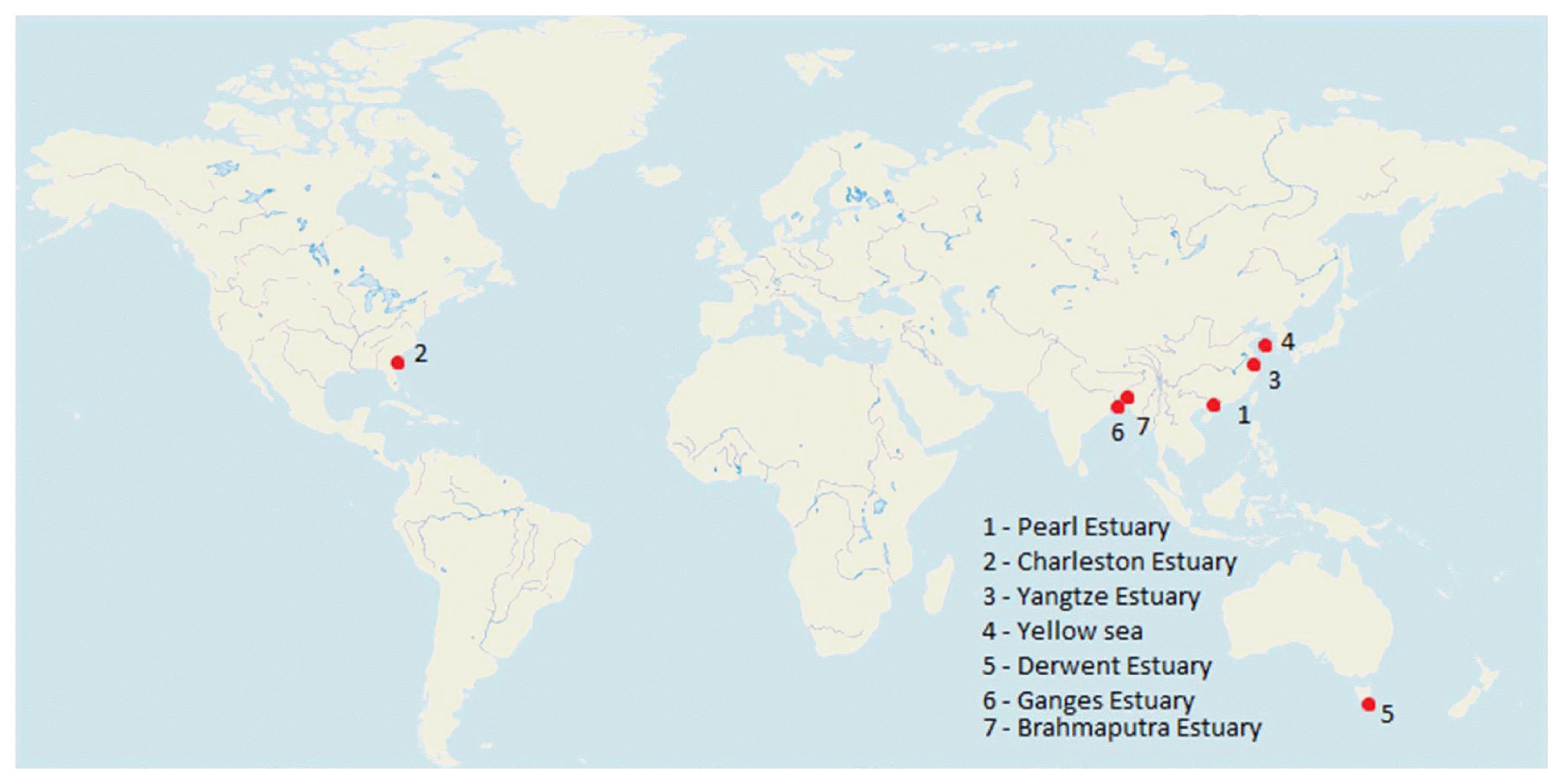
Fig. 3Schematic showing sources of antibiotic resistant genes and horizontal gene transfer within biofilm developed on microplstics in estuaries. 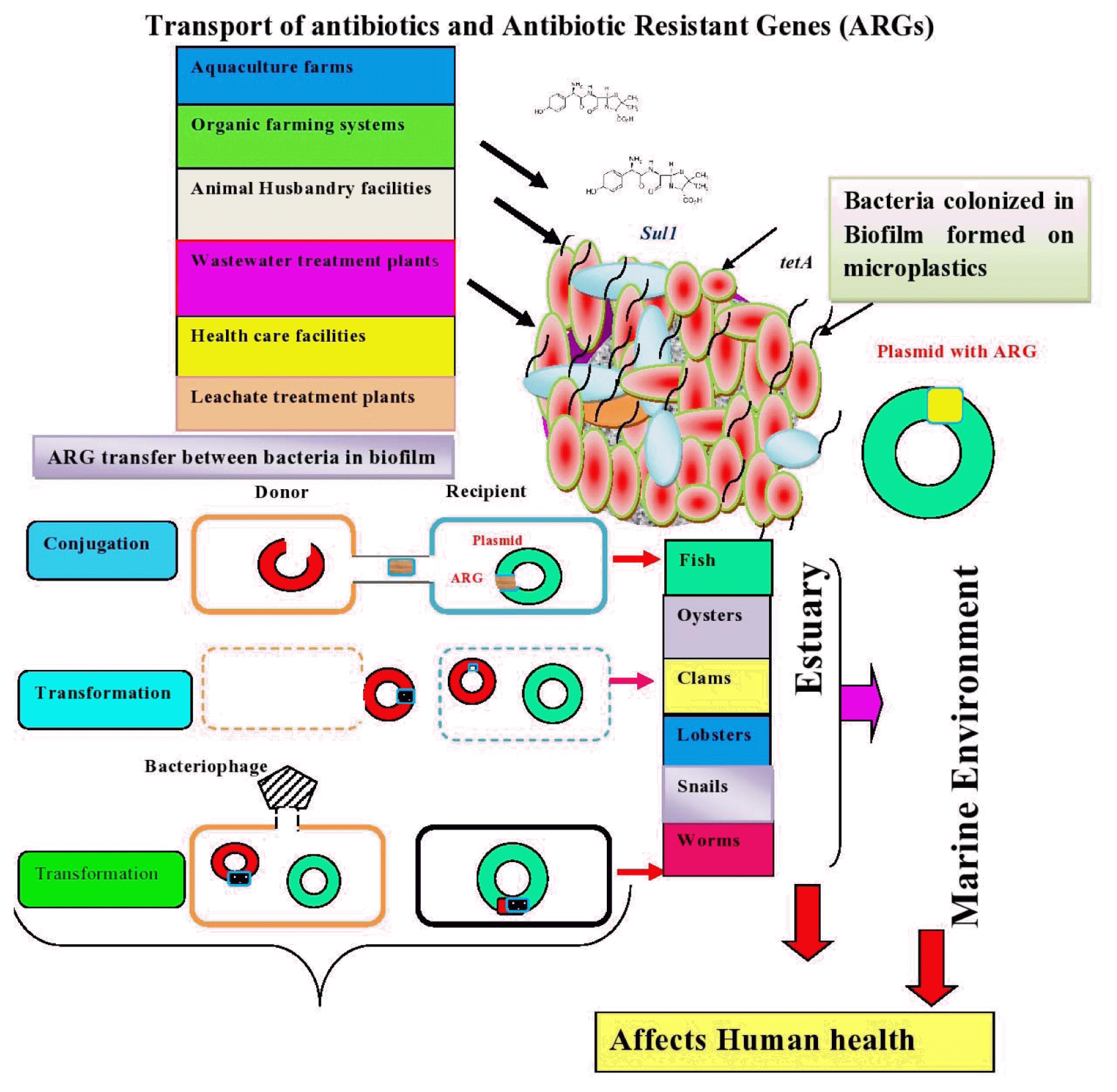
Table 1Occurrence of different sizes, forms, and types of Microplastics in Estuaries across the world
NS - Not studied; RY – Rayon; PS - Polystyrene; PA - Polyamide; PES – Polyester; PVC - polyvinyl chloride; PET - Polyethylene Terephthalate; AC - Acrylic; PAM - Polyacrylamide; PEVA - Polyethylene vinyl acetate; PAN – Polyacrylonitrile; LDEPE - Low density polyethylene; HDPE - High density polyethylene; PP - Polypropylene; PVP - Polyvinylpyrrolidone; EVA - Polyethylene covinyl acetate Table 2Occurrence of Antibiotic Resistant Genes in Estuaries
Multi Drug Resistant P. aeruginosa has been shown to form strong to weak biofilms. P. aeruginosa biofilm developed increased resistance to piperacilin, cefepime, aztreonam, imipenem, and mesopenem, which were reported as strong biofilm formation. In addition to the MexAB-OprV efflux pump system, three other efflux pump systems, MexCD-OprJ, MexXY-OPrM, and MexEF-OprN, contribute to antibiotic resistance [138]. Alternately, bacteria that exhibit antibiotic resistance as a result of the bacterial cells’ exposure to certain environmental stress, which enables bacteria to respond more quickly to an antibiotic challenge, typically reverts to the original form once the inciting signal is eliminated [139]. Table 3Predominant antibiotic resistance genes occur in estuaries, their characteristics and sources
Table 4Bacterial Community Structure of Biofilms Formed on MPs in the Estuary
Table 5Important factors affecting adsorption of microorganisms on MPs during biofilm formation
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||