AbstractFree cyanide (CN−) is a threat to metabolic functions of the microbial population used for the treatment of wastewater, particularly, total nitrogen removal (TN) consortia which gets inhibited by CN− in wastewater treatment plants (WWTPs). Many other methods are used to treat CN− prior to the TN removal stages; however, these methods increase the operational cost of the WWTPs. The capability of a microbial population to use multiple substrates is critical in WWTP and in eliminating inhibition associated with CN−. Previously, cyanide resistant bacteria were used to eliminate the inhibitory effect of CN− towards simultaneous nitrification and aerobic denitrification (SNaD). However, a study to predict the degradation efficiency of the microorganism was required. In this study, response surface methodology (RSM) and cybernetic models were used to predict and optimize SNaD performance for TN removal under CN− conditions. Physiological parameters influencing the SNaD were pH 6.5 and 36.5°C, with TN and CN− degradation efficiency of 78.6 and 80.2%, respectively. These results show a complete elimination of the CN− inhibitory effect towards SNaD and show the prediction ability of both RSM and the cybernetic models used. These results exhibited a promising solution in the control, management, and optimization of SNaD.
1. IntroductionThe growth of a microorganism under multiple carbon source conditions can influence its performance even in large-scale wastewater treatment plants (WWTPs) [1–3]. Similarly, wastewater can be composed of different nitrogenous compounds from several industries. Current methods to treat this type of wastewater consist of traditional two-stage biological processes known as aerobic nitrification and anoxic denitrification. These treatment processes have different operating conditions and requirements such as dissolved oxygen concentration, carbon source requirements, and retention time, with throughput rates of the wastewater being treated in the processes being of utmost importance. Due to high energy consumption associated with the second reactor dedicated to anoxic denitrification, some studies have indicated the possibility of simultaneous nitrification and aerobic denitrification (SNaD) [4]. This process is advantageous than traditional nitrification after denitrification, providing for cost-effectiveness [5–7]. Moreover, SNaD is a highly effective method for total nitrogen (TN), i.e. ammonium nitrogen (NH4-N), nitrate-nitrogen (NO3-N), and nitrite-nitrogen (NO2-N), removal from wastewater [8]. However, the microorganism(s) used during TN removal are highly sensitive to toxicant loadings, particularly free-cyanide (CN−), thus, SNaD is the least effective of industrial wastewater treatment methods owing to a high concentration of contaminants that may completely inhibit SNaD [9–11]. Some remedial strategies such as activated carbon and chemical treatment have been used as a pretreatment step to combat the effect of CN− on TN removal. Similarly, activated carbon has been reported to have low adsorption capability towards CN− [12]. Furthermore, this additional step increases the operational cost of WWTPs associated with the operation of an additional reactor dedicated to the pretreatment of CN−. Thus, biological treatment methods, are recommended as a sustainable strategy to resolve the inhibitory effects of CN− in TN removal [12].
The type of wastewater containing CN− may result in serious environmental contamination and other challenges associated with its disposal if not treated [13–15]. Kim et al. [5] have reported that CN− possess the highest inhibitory effect on SNaD. Previously, CN− inhibition has been eradicated by biological, physical, and chemical pretreatment methods [16]. These methods have been reported to increase operational costs associated with the operation of multiple reactors for pretreatment of the wastewater before SNaD. Hence, some studies have proposed the application of cyanide degrading bacteria for SNaD to eliminate the inhibitory effect of CN− towards SNaD, which can further reduce operational costs associated with the reactor designated for biological pretreatment of CN− [17] and for downstream denitrification.
Mpongwana et al. [18] and Han et al. [19] recommended the application of CN− degrading bacteria for TN removal from wastewater containing a high quantity of CN− as an alternative to mediate the effect of CN− on SNaD. Other researchers have also recommended this strategy as a sustainable solution to SNaD inhibition by CN− [20, 21]. Moreover, the application of cyanide degrading bacteria will not only resolve the inhibitory effect of CN− towards SNaD, it will also resolve the challenge of slow-growing microorganisms responsible for SNaD since cyanide degrading bacteria have higher growth rates [22, 23]. This will likely improve process performance and compensate for biomass washout. Also, some cyanide degrading bacteria have been reported to also possess other important genes that are responsible for polyhydroxyalkanoates (PHA) synthesis which can serve as a carbon source during the cyanide degradation process [24]. This is another advantageous trait of cyanide degrading bacteria.
Mpongwana et al. [25], also reported a complete eradication of CN− inhibition towards SNaD by using a cyanide resistant A. courvalinii; however, the presence of CN− reduces the degradation rate of TN. Thus, this study aims to optimize TN removal by A. courvalinii under CN− conditions. The presence of CN− compound alters the metabolic functions of microbial populations [5]; however, modelling approaches used to model single staged uninhibited SNaD systems may not adequately represent SNaD as a process on an industrial scale, particularly when potent inhibitors such as CN− are readily prevalent in TN containing wastewater. Hence, it is imperative to study the predictive ability of response surface methodology (RSM) which is a commonly used optimization and performance predictive software including a metabolic network model which also considers the metabolic functions of the microbial population since CN−/SNaD is an enzyme facilitated process.
2. Materials and MethodsThe methodological flowchart for this study is illustrated in Fig. 1. The first step involved the isolation of different bacterial strains (n = 16) from CN− containing wastewater, followed by determining their resistance to varying concentrations of CN− and NH4NO3 [26]. Some strains (n = 4) were found to be able to grow under high concentrations of CN− and ammonium. However, a single strain was found to be able to grow and utilize both CN− and TN as nitrogenous sources in a simulated SaND process. RSM was then used to determine the optimum conditions, with further reductions to the optimum quadratic model generated being implemented to adequately describe the process under study. Subsequently, CN− and TN degradation kinetics experiments ensued using the optimum conditions obtained from the RSM, followed by cytoplasm extraction for application in enzyme kinetics. Similarly, a cybernetic model was developed and used to predict the rate of CN− and TN degradation as well as enzyme kinetics. After the conclusion of these experimental steps, the process prediction using models developed were regressed using polymath 6.0.
2.1. Microbial Isolation and IdentificationThe bacteria used for this study was isolated from CN− containing waste at the Bioresource Engineering Research Group (BioERG) facility at the Cape Peninsula University of Technology (CPUT), South Africa. Isolates were cultured on nutrient agar to obtain pure colonies. Thereafter, they were grown on nutrient agar (NA) containing different concentrations ranging from 10 to 300 mg CN−/L to determine the highest concentration of CN− tolerance, after gram staining. The selected cyanide tolerant bacteria’s 16S RNA was sequenced and the bacteria was identified as A. courvalinii (accession number AB602910.1 or NR_148843.1) [27].
2.2. Response Surface Methodology2.2.1. Central composite design experimentsThe central composite design was used for the optimization of SNaD under CN− conditions. This was done by determining the independents’ variables, i.e. temperature and pH, optima which have been reported to affect SNaD [28]. In this study, a 13-run experimental plan which included the variation in independent variables (temperature and pH) was generated using central composite design (CCD) (see Table S1).
The experimental design used for this study is shown in Table S2, with the corresponding response being primarily TN removal and CN− degradation. Synthetic wastewater containing 20 mg NH4-N /L and 20 mg CN− /L were used for these experiments. The isolated strain was grown in a 250 mL multiport Erlenmeyer flask with 40 mL basal media containing: 1.5 g KH2PO4, 7.9 g Na2HPO4, 0.5 g MgSO4.7H2O and 1 mL of trace elemental solution per litre. The trace element solution contained (per litre): 50 g EDTA, 2.2 g ZnSO4.7H2O, 5.5 g CaCl2, 5.06 g MnCl2.4H2O, 5.0 g FeSO4.7H2O, 1.1 g (NH4)6Mo7O2.4H2O, 1.57 g CuSO4.5H2O, 1.61 g CoCl2.6H2O. The isolate was allowed to grow for 24 h before the addition of toxicants; KCN as CN− and NH4SO4 as NH4-N.
The Erlenmeyer flasks that were used for this experiment had a sealable sampling port to avoid volatilization of CN−. After the addition of the toxicant, each experiment was monitored for 5 h, and samples were taken after every 1 h to analyse for CN−, and TN as NH4-N, NO2-N, and NO3-N, using Merck test kits and a Merck Spectroquant. All the experiments were done in duplicates with control experiments that did not contain the microorganism to account for the volatilization of CN− and NH4-N stripping. To determine the critical points (maximum, minimum, target, or within range) a polynomial function that contains quadratic terms were used (Eq. (1)) [29].
Where k is the number of variables, β0 is the constant term, βi is the coefficients of the linear parameters, βij is the coefficients of the interaction parameters, βii is the coefficients of the quadratic parameter, xi represents the variables, and ɛ is the residual associated with the experiments.
2.2.2. Data handlingAll the experiments were conducted in duplicates and the mean were calculated according to Eq. (2), thereafter the data was computed into Microsoft Excel 2016.
Where ∑X is the sum of all the duplicated data points. While n is the number of data points. Standard deviation and variance were obtained from RSM and Polymath respectively. Standard deviation was calculated from variance using Eq. (3).
Where s and σ are standard deviation and variance, respectively
2.2.3. Statistical analysesThe statistical analyses were centred on the lack of fit test which describes the fitness of mean and reduced the quadratic model of TN removal and the sequential model sum of squares. The model as represented in Eq. (1) indicated the relationship between the dependent variables and TN removal efficiency by the isolate in CN− containing wastewater. Furthermore, the significance of each variable in the model was analysed using Analysis of Variance (ANOVA) with a Multiple Regression Analysis being used to analyse the experimental data obtained.
2.3. Cybernetic Model2.3.1. Batch culture experimentA basal media similar to that used for the RSM experiments was used in batch cultures; although, the experiment was carried out in 1 L reactors. The media was inoculated with 100 mL of an overnight grown culture (24 h) incubated at optimum conditions determined by RSM, i.e. 36.5°C, and a media pH of 6.5. The culture was incubated for 168 h before the supplementation of 40 mg CN−/L and 250 mg NH4-N/L; a combination that was shown to have little growth inhibition during toxicity assessments, with 2 mL samples being taken periodically (24 h intervals) for analyses, i.e. CN−, NH4-N, NO2-N, and NO3-N. Enzymes, assumed to have been produced extracellular, were extracted using cold acetone daily from the cultures, to quantify maximum enzyme activity (
).
2.3.2. Enzyme activity assessmentsThe suspended bacterial cells in samples were removed by centrifuging each sample at 5,000 g for 5 min; thereafter, to the cell-free supernatant, cold acetone was added, forming a cytoplasm precipitate, herein referred to as the enzyme extract, which was then separated from the supernatant by centrifugation at 16,000 g for 15 min. The precipitate was washed (n = 3) and initially stored at −18 °C prior to re-suspension in 360 mg/L phosphate buffer solution (pH 7.4). NH4-N and CN− solutions with an initial concentration of 10 mg/L for each contaminant were prepared and the enzyme extract was added into the solution, while the changes in CN− (09701), NH4-N (00683), NO2-N (110057), and NO3-N (14773) were monitored throughout the experiment.
2.3.3. Analytical proceduresAll the test kits used for the analysis of the samples were obtained from Merck SA (Merck Pty Ltd, Modderfontein, South Africa). Furthermore, a Merck Spectroquant Nova 60 instrument was used to analyse residual CN−, NH4-N, NO2-N, and NO3-N. The basis of the individual assays is described elsewhere [30]. The kits were used as per the manufacturer’s instructions.
2.3.4. Model developmentA simplified model for the prediction of SNaD as well as CN− degradation in a single reactor was developed as shown in Fig. 2.
Two nitrogenous compounds NH4-N and CN− were used as pollutants with S1 and S2 presenting NH4-N which is assimilated none-enzymatically into the cell and CN− respectively, while M1, M2, and M3, represented intermediates, NH4-N, NO2-N, and NO3-N respectively. The biomass used was deemed capable of using both NH4-N and CN− as primary pollutants to be degraded. The biomass converts CN− into NH4-N for which a part of NH4-N is assimilated for proliferation [31] and the other portion is further converted into intermediates NO2-N and NO3-N.
The presence of NO3-N can induce denitrification enzymes production thus resulting in the initiation of the denitrification process which involves the conversion of NO3-N denoted as M3 into nitrogenous gas (N2) denoted as P1. Under optimized conditions, it has been shown that when the biomass is provided with NH4-N and CN− it undergoes multiphase growth while simultaneously degrading the pollutants. Therefore, the predictive capability of cybernetic models was evaluated to estimate the simultaneous degradation of NH4-N and CN− using the cyanide degrading bacteria A. courvalinii. The model was developed based on the metabolic reaction network as illustrated in Fig. 1.
All these processes are catalysed by specific enzymes, i.e. ammonia monooxygenase (AMO), nitrate reductase (NaR), and nitrite reductase (NiR). When modelling metabolic networks, two vectors are considered important for cybernetic variables. These vectors are and v, for which is a fractional allocation of resources needed for enzyme synthesis, such that
and with v representing the activity of the different enzymes. Conditions 0 ≤ ui, vi ≤ 1, whereby. i = 1,2,3,4. u is needed for balancing the reaction. For example, when the maximum synthesis rate rE for enzyme Ei is regulated then ui = rEi. Moreover, vi is required to estimate the regulated reaction rates; thus, ri will be the rate of the ith reaction when enzyme Ei is fully active. The regulated ith reaction can also be written as r = vrri.
Monods’ model was used to express the rate of TN removal (ri) for pollutant Si and Sj catalysed by enzyme ei to form intermediate M2 and M3 (Eq. (4)).
The cell growth rate (rg) was model using rate law equation (1st order) as indicated in Eq. (5).
Where
is the are maximum pollutant removal rate and Ki is the pollutant saturation constant, while X is cell concentration and μg is the cell-specific growth rate.
Enzyme synthesis for the two pollutants, S1 and S2 was assumed to be maximized as the growth rate of biomass increased such that (Eq. (6));
It was hypothesised that NH4-N was broken down into NO2-N via AMO (represented by e2) catalysis while CN− was decomposed by cyanide degrading enzyme(s) represented as e1. The intermediate NO2-N was further broken down into NO3-N facilitated by NiR represented in the diagram as e3, with NO3-N being further decomposed into N2 by NaR represented as e4. The model, Eq. (7) would thus represent the rate of enzyme(s) synthesis responsible for the SNaD under cyanogenic conditions.
Where
is the rate of enzyme(s) synthesis, while αei represents an inductive rate, bi the activity of the enzyme responsible for decomposition of a pollutant, rei the decomposition rate of the nitrogenous pollutant, bi is the degradation rate constant of the enzyme, Dg dilution term due to growth rate. All these model parameters were estimated by polymath software v6.0 using experimental data.
3. Results and Discussion3.1. Predictive Ability of Response Surface Methodology3.1.1. Analysis of variance (ANOVA) for TN removalThe central composite design was used to study the interaction between the independent variables, pH and temperature, that affect SNaD for TN removal. Table 1 shows analysis of variance (ANOVA) of the quadratic model used to describe TN removal. The model F-value of 17.01 for TN removal was higher than 1.0, which indicated that the variation between the model and experimental data was higher.
The predicted values of the RSM were compared with the experimental values, with the p-value being 0.0009 for the model, which is smaller than the alpha level of 0.05. This meant that there was a consequential relationship between the predicted values and the actual values of TN removal. However, other parameters such as the deviation of the model values from the actual data points (standard deviation), R2, adjusted R2, and predicted R2 need to be considered to judge the adequacy of the model – see Table 1.
The quadratic model was also used to predict SNaD under CN− conditions. The significance of the parameters was determined using values of “Prob > F” less than 0.05, thus, in this case, A, A2, and B2 were found to be significant for the model. Therefore, the model was improved by reducing it from Eq. (8) to Eq. (9).
The visualization of the predicted model was obtained using a surface response plot – see Fig. 3. In this study, the response was set to maximize the degradation efficiency of TN in a SNaD system that contains multiple nitrogenous sources, i.e NH4-N, NO2-N, and NO3-N, with CN− as a secondary nitrogenous source or pollutant. A. courvalinii was found to remove up to 78.6% of TN via SNaD from an initial concentration of 20 mg NH4-N/L within 5 h of incubation. Li et al. [32] reported that Pseudomonas stutzeri YZN-001 could remove NH4-N at 37 °C rapidly; however, the removal of NO2-N and NO3-N only occurred at 30°C.
The surface plot for TN removal under cyanogenic conditions as shown in Fig. 3, indicated that the maximum operational efficiency for the SNaD containing CN− was located inside the experimental region. This was a confirmation of the optimum being 6.5 and 36.5°C. Moreover, the plateau in surface response plots in Fig. 3 indicated the suitability of operational conditions for SNaD, it also highlighted that the lowest degradation efficiency for TN removal and CN− degradation was observed at temperature and pH of 40.04°C and 6.5 with degradation efficiencies of approximately 5 %. When the temperature is 36.5 °C with pH being 2.9 or 10.0 the degradation efficiency for TN removal was found to be 46.1 and 5.6%, respectively. This indicated that changes in pH and temperature affect TN removal.
3.1.2. Batch reactor experiment and model simulationsTo generate data for the models, removal (degradation) kinetics of TN were studied at optimized conditions in 1L reactors. The initial (higher) concentration of 40 mg CN−/L and 250 mg NH4-N/L were used and the experiment was conducted for 264 h. The results indicated that up to 70.5% of TN was removed within 264 h, see Fig. 4(a). A. courvalinii was shown to possess distinctly different attributes involving the simultaneous removal/ or degradation of TN and CN−, which is an indication of the complete elimination of inhibitory effects of CN− compound towards SNaD used for TN reduction in wastewaters. Duan et al. [33] reported that a nitrification efficiency of 91.82 ± 1.98% after 42 h by Vibrio diabolicus SF16 was achievable in a system that does not contain CN−.
Furthermore, He et al. [10] also reported an NH4-N removal efficiency of 93.6% after 96 h of incubation with Pseudomonas tolaasii Y-11; albeit, the utilization of pollutants by a specific species might be sequential, with the organism utilizing the easily biodegradable pollutant with a less complex structure first, subsequent to the degradation of the second less desirable pollutants which sometimes results in multi-phased growth of the organism(s) used. The cell concentration was also studied and modelled (Fig. 4(a)).
The data points of predicted growth rate versus actual growth rate were scattered, with numerous outliers from the trend line (Fig. 4(b)) which signifies that the first-order equation did not adequately represent the growth model with the variance being 4.75 × 109 and a standard deviation of 6.89 ×104 and low R2 of 0.5303 indicating a poor prediction of the growth. Song and Lui [34] also reported that simulation of cybernetic model adequately matches with the biomass proliferation profile of Acidovorax sp. and P. denitrificans. This lack of fit was associated with other factors that were not considered by the model such as the inhibition of the microbial growth by metabolic by-products. Overall, the microorganism used in this study primarily degraded TN including CN− simultaneously, which is an interesting trait considering that CN− is a known inhibitor to nitrification, with as little as 1 mg CN−/L completely inhibiting nitrification [5].
3.2. Prediction Ability of RSM in Comparison to Cybernetic ModelsWhen comparing modeled and actual SNaD performance under optimum conditions, the correlation coefficient (R2) of 0.92 was observed; although this high R2 was observed, R2 alone cannot verify whether the model is adequate or not, since R2 can improve with the extension of dependent variables scale regardless of whether the variable is significant or not. Thus the adjusted R2 (0.87) and predicted R2 (0.46) were considered (Table S3). Adjusted R2 is normally used to compare the explanatory power of a regression model, while the Adjusted R2 only account for variables that improve the model; therefore, the addition of less significant variables decreases the Adjusted R2. Overall, the Adjusted R2 is considered more reliable compared to the correlation coefficient.
Hence, the Adjusted R2 was used to evaluate the adequacy of the model culminating in the selection of a suitable model describing SNaD for TN removal under CN− conditions. As the Adjusted R2 of 0.87 was higher; therefore, the model was deemed adequate. However, the difference between predicted R2 and adjusted R2 was above 0.2, indicating that there would be challenges with the use of the model. Hence, the model was reduced to expression as highlighted in Eq. (7).
This was further confirmed by the average standard deviation (12.11) for TN removal with a standard error (SE) of 5.284 (Table S3). The SE for TN removal indicated that the RSM model could be improved to represent the experimental data more adequately. The parity plot for TN removal was used to compare the predicted TN removal efficacy and the actual TN removal efficiency. The data points of the RSM model deviated from the trendline indicating a larger deviation of the data points from the trend line.
Cybernetic models were developed to predict maximum pollutant utilization rates which are presented as S1 and S2 for TN and CN−, respectively via catalysis by numerous enzymes. A cybernetic model was simulated by estimating unknown parameters using experimental data as listed in Table 2. The model successfully described the rate of TN removal with (R2 of 0.97, indicating a 97% suitability. Moreover, the Adjusted determination coefficient (Adj R2 = 0.96) was very high with the difference between R2 and the Adj R2 being 0.006. This difference is minute; therefore, it advocated for the high significance of the model.
Furthermore, the variance (0.012), standard error (0.0035) and standard deviation (0.047) for TN removal were low, demonstrating that there’s an insignificant difference between the predicted values from the model and the actual experimental values. Although the RSM model was shown to adequately represent TN removal, the adjusted R2 of the cybernetic model was higher than that of the RSM model. Thus, cybernetic models were selected as the best model to predict TN removal using SNaD. Song and Liu. [34] also reported a better prediction of denitrification by a cybernetic model compared to a simple kinetic model. This proves that biological processes such as TN removal must be predicted using metabolic network modelling that takes into account the metabolic functions of the microbial population involved in the process. Furthermore, the parity plot for RSM and the cybernetic model were compared. The parity plot of the cybernetic model had data points scattered closer to the trend line as opposed to the RSM model which numerous outliers showing a significant deviation from the trend line (Fig. 5(b) and (d)), supporting the statement that the cybernetic model better-predicted TN removal better in CN− using SNaD.
3.3. TN/ CN- BiocatalysisSNaD involves a process whereby NH4-N is converted into NO2-N and further to NO3-N which activates the production of denitrification enzymes that convert NO3-N into N2. Enzyme activity was determined by extracting free-cell enzymes supplemented into solutions consisting of pollutants, NH4-N, and CN−. The decrease in the pollutants and the accumulation of the intermediates NO3-N and NO2-N were observed (Fig 6(a)–(f)). The decrease in NH4-N and CN− was slow in the first two minutes of the reaction with a degradation rate of 0.83 and 1.1 mg/L/min, respectively.
The degradation rates increased after the fourth minute, with the rate increasing up to 1.64 and 1.73 mg/L/min being observed, respectively; albeit, the rate of degradation decreased after 5 min. Since the degradation was conducted at ambient temperature to simulate real-life WWTP conditions, the decrease in the pollutants indicated the presence of TN and CN− degrading enzymes; moreover, the increase and decrease of intermediates NO2-N and NO3-N indicated the presence of denitrification enzymes; hence, the isolated microorganisms were deemed to be capable to carry-out SNaD even in the presence of CN−. The enzyme synthesis model successfully described individual enzyme activity with R2 above 0.7 indicating a good fit of the activity models. Hamilton et al [35] also reported that enzyme biosynthesis based models are better predictors of microbial growth, which supports the results obtained from this study whereby it was shown that the cybernetic model is a better predictor of multiple nitrogenous source utilization such as CN− and TN in systems such as SNaD.
3.4. Overall Remarks about the StudyThe results obtained from this study indicated that the inhibition effect of CN− towards SNaD can be completely eradicated by using cyanide degrading microorganism(s) for SNaD systems, provided that appropriate operational conditions are established. This eliminates the necessity of using a pre-treatment reactor for CN− removal, thus, lowering operational costs. Additionally, these findings provide a sustainable solution towards inhibition reduction of SNaD in large-scale WWTPs. The RSM and cybernetic models were compared for their ability to predict SNaD performance even in the presence of inhibitors such as CN−, which is known as a highly potent toxicant. The findings indicated that the cybernetic modeling adapted from metabolic networking provides an approach that is a robust and novel predictor of SNaD under inhibitory conditions, even when novel hybridized models, useful for process control of SNaD, are used.
4. ConclusionResponse surface methodology was used to determine the optimum conditions for TN removal under CN− conditions using A. courvalinii, a CN− tolerant bacterium with TN removal capabilities in SNaD. The optimum pH and temperature were found to be 6.5 and 36.5°C, with degradation efficiency of 78.6% for TN and a significant (80.2%) for CN− degradation, respectively. Moreover, cybernetic modeling was found to be better at predicting SNaD under cyanogenic conditions with a higher adjusted R2 (0.96) as opposed to the RSM models with an adjusted R2 of 0.87. The cybernetic models used in this study were simplified; thus they do not provide illuminating insights into cellular responses. Hence, additional experiments are needed to elucidate metabolic flux distributions better and to further develop models for SNaD process prediction as well as optimisation.
AcknowledgmentsThis study was supported by the University Research Fund (URF RK16) at the Cape Peninsula University of Technology.
NotesAuthor Contributions N.M. (Ph.D) conducted all the experiment and wrote the paper. E.I.O. (Ph.D) interpreted data and revised the manuscript. L.C.R. (Ph.D) interpreted data and revised the manuscript. B.S.C. (Ph.D) interpreted data and revised the manuscript. S.K.O.N. (Associate Professor) interpreted data, acquired funding and editing the manuscript. References1. Van Dedem G, Moo-Young MA. Model for diauxic growth. Biotechnol Bioeng. 1975;17:1301–1312.
2. Narang A, Pilyugin S. Bacterial gene regulation in diauxic and non-diauxic growth. J Theor Biol. 2007;244:326–348.
3. Solopova A, van Gestel J, Weissing F, et al. Bet-hedging during bacterial diauxic shift. Proc Natl Acad Sci USA. 2014;111:7427–7432.
4. Chen P, Li J, Li QX, et al. Simultaneous heterotrophic nitrification and aerobic denitrification by bacterium Rhodococcus sp. CPZ24. Bioresour Technol. 2012;116:266–270.
5. Kim Y, Park D, Lee D, Park J. Inhibitory effects of toxic compounds on nitrification process for cokes wastewater treatment. J Hazard Mater. 2008;152:915–921.
6. Zhang Y, Shi Z, Chen M, Dong X, Zhou J. Evaluation of simultaneous nitrification and denitrification under controlled conditions by an aerobic denitrifier culture. Bioresour Technol. 2015;175:602–605.
7. Jin R, Liu T, Liu G, Zhou J, Huang J, Wang A. Simultaneous heterotrophic nitrification and aerobic denitrification by the marine origin bacterium Pseudomonas sp ADN-42. Appl Biochem Biotechnol. 2015;175:2000–2011.
8. He T, Li Z, Sun Q, Xu Y, Ye Q. Heterotrophic nitrification and aerobic denitrification by Pseudomonas tolaasii Y-11 without nitrite accumulation during nitrogen conversion. Bioresour Technol. 2016;200:493–499.
9. Choi OK, Hu ZQ. Nitrification inhibition by silver nanoparticles. Water Sci Technol. 2009;59:1699–1702.
10. Papirio S, Zou G, Ylinen A, Di Capua F, Pirozzi F, Puhakka JA. Effect of arsenic on nitrification of simulated mining water. Bioresour Technol. 2014;164:149–154.
11. Ma Q, Qu Y, Shen W, et al. Bacterial community compositions of coking wastewater treatment plants in steel industry revealed by Illumina high-throughput sequencing. Bioresour Technol. 2015;179:436–443.
12. Singh N, Balomajumder C. Simultaneous removal of phenol and cyanide from aqueous solution by adsorption onto surface modified activated carbon prepared from coconut shell. J Water Process Eng. 2016;9:233–245.
13. Feng C, Huang L, Yu H, Yi X, Wei C. Simultaneous phenol removal, nitrification and denitrification using microbial fuel cell technology. Water Res. 2015;76:160–170.
14. Carrera J, Baeza J, Vicent T, Lafuente J. Biological nitrogen removal of high-strength ammonium industrial wastewater with two-sludge system. Water Res. 2003;37:4211–4221.
15. Kim Y, Cho H, Lee D, Park D, Park J. Comparative study of free cyanide inhibition on nitrification and denitrification in batch and continuous flow systems. Desalination. 2011a;279:439–444.
16. Kim Y, Lee D, Park C, Park D, Park J. Effects of free cyanide on microbial communities and biological carbon and nitrogen removal performance in the industrial activated sludge process. Water Res. 2011b;45:1267–1279.
17. Cui J, Wang X, Yuan Y, Guo X, Gu X, Jian L. Combined ozone oxidation and biological aerated filter processes for treatment of cyanide containing electroplating wastewater. Chem Eng. 2014;241:184–189.
18. Han Y, Jin X, Wang Y, Liu Y, Chen X. Inhibitory effect of cyanide on nitrification process and its eliminating method in a suspended activated sludge process. Environ Sci Pollut Res Int. 2014;21:2706–2713.
19. Mpongwana N, Ntwampe SKO, Omodanisi EI, Chidi BS, Razanamahandry LC. Sustainable approach to eradicate the inhibitory effect of free-cyanide on simultaneous nitrification and aerobic denitrification during wastewater treatment. Sustainability. 2019;11:6180.
20. Han Y, Jin X, Wang F, Liu Y, Chen X. Successful startup of a full-scale acrylonitrile wastewater biological treatment plant (ACN-WWTP) by eliminating the inhibitory effects of toxic compounds on nitrification. Water Sci Technol. 2013;69:553–559.
21. Park D, Lee D, Kim Y, Park J. Bioaugmentation of cyanide-degrading microorganisms in a full-scale cokes wastewater treatment facility. Bioresour Technol. 2008;99:2092–2096.
22. Han Y, Jin X, Wang Y, Liu Y, Chen X. Inhibitory effect of cyanide on nitrification process and its eliminating method in a suspended activated sludge process. Environ Sci Pollut Res. 2013;21:2706–2713.
23. Ojaghi A, Tonkaboni SZS, Shariati P, Ardejani FD. Novel cyanide electro-biodegradation using Bacillus pumilus ATCC 7061 in aqueous solution. J Environ Health Sci eng. 2018;16:99–108.
24. Kandasamy S, Dananjeyan B, Krishnamurthy K, Benckiser G. Aerobic cyanide degradation by bacterial isolates from cassava factory wastewater. Braz J Microbiol. 2015;46:659–666.
25. Luque-Almagro VM, Moreno-Vivián C, Roldán MD. Biodegradation of cyanide wastes from mining and jewellery industries. Curr Opin Biotechnol. 2016;38:9–13.
26. Mpongwana N, Ntwampe S, Mekuto L, Akinpelu E, Dyantyi S, Mpentshu Y. Isolation of high-salinity-tolerant bacterial strains, Enterobacter sp, Serratia sp, Yersinia sp, for nitrification and aerobic denitrification under cyanogenic conditions. Water Sci Technol. 2016;73:2168–75179.
27. Mpongwana N. Metabolic network modelling of nitrification and denitrification under cyanogenic conditions [dissertation]. South Africa: Cape Peninsula University of Technology; 2019.
http://hdl.handle.net/20.500.11838/2982
28. Stephen FA, Thomas LM, Alejandro AS, et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–3402.
29. Mekuto L, Ntwampe SKO, Jackson VA. Biodegradation of free cyanide and subsequent utilisation of biodegradation by-products by Bacillus consortia: optimisation using response surface methodology. Environ Sci Pollut Res Int. 2015;22:10434–10443.
30. Mpongwana N, Ntwampe SK, Mekuto L, Akinpelu EA, Dyantyi S, Mpentshu Y. Isolation of high-salinity-tolerant bacterial strains, Enterobacter sp., Serratia sp., Yersinia sp., for nitrification and aerobic denitrification under cyanogenic conditions. Water Sci Technol. 2016;73:2168–2175.
31. Kanyenda G, Ntwampe SKO, Mpongwana N, Godongwana B. Mathematical exposition of simultaneous nitrification and aerobic denitrification. In : Proceedings of the 10th lnt’I Conference on Advances in Science, Engineering, Technology & Healthcare (ASETH-18); 19–20 November 2018; Cape Town, South Africa. p. 19–20.
32. Li C, Yang J, Wang X, et al. Removal of nitrogen by heterotrophic nitrification–aerobic denitrification of a phosphate accumulating bacterium Pseudomonas stutzeri YG-24. Bioresour Technol. 2015;182:18–25.
33. Duan J, Fang H, Su B, Chen J, Lin J. Characterization of a halophilic heterotrophic nitrification–aerobic denitrification bacterium and its application on the treatment of saline wastewater. Bioresour Technol. 2015;179:421–428.
34. Song HS, Liu C. Dynamic metabolic modelling of denitrifying bacterial growth: the cybernetic approach. Ind Eng Chem Res. 2015;54:10221–10227.
Fig. 2Simplified metabolic network diagram of SNaD under cyanide-laden conditions. Key: S1: NH4-N, S2: CN−, M1: NH4-N, M2: NO2-N, M3: NO3-N, and P1: N2.
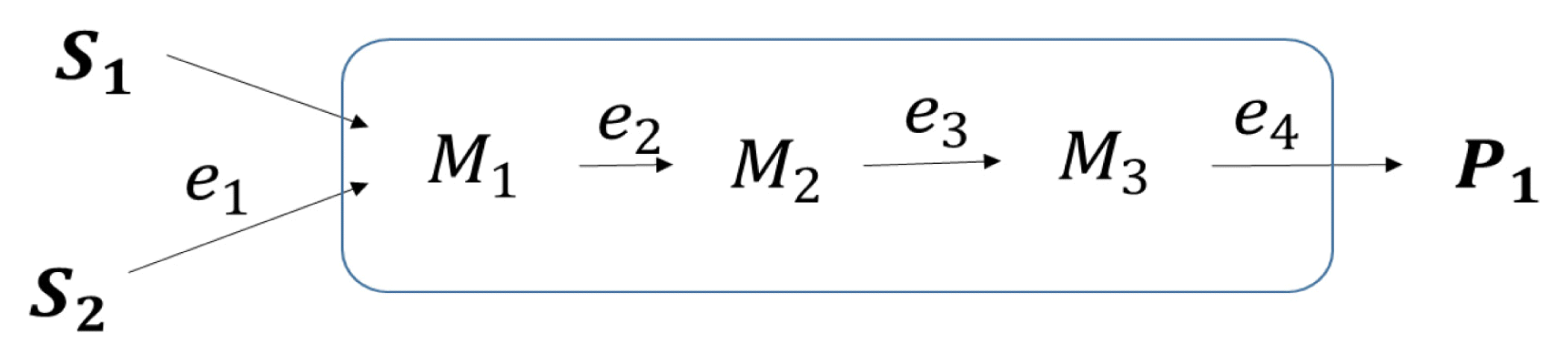
Fig. 4Degradation kinetics of TN and cyanide in a batch culture reactor. A: TN and CN− degradation and cell concentration over time. B: model fitting into biomass plot. 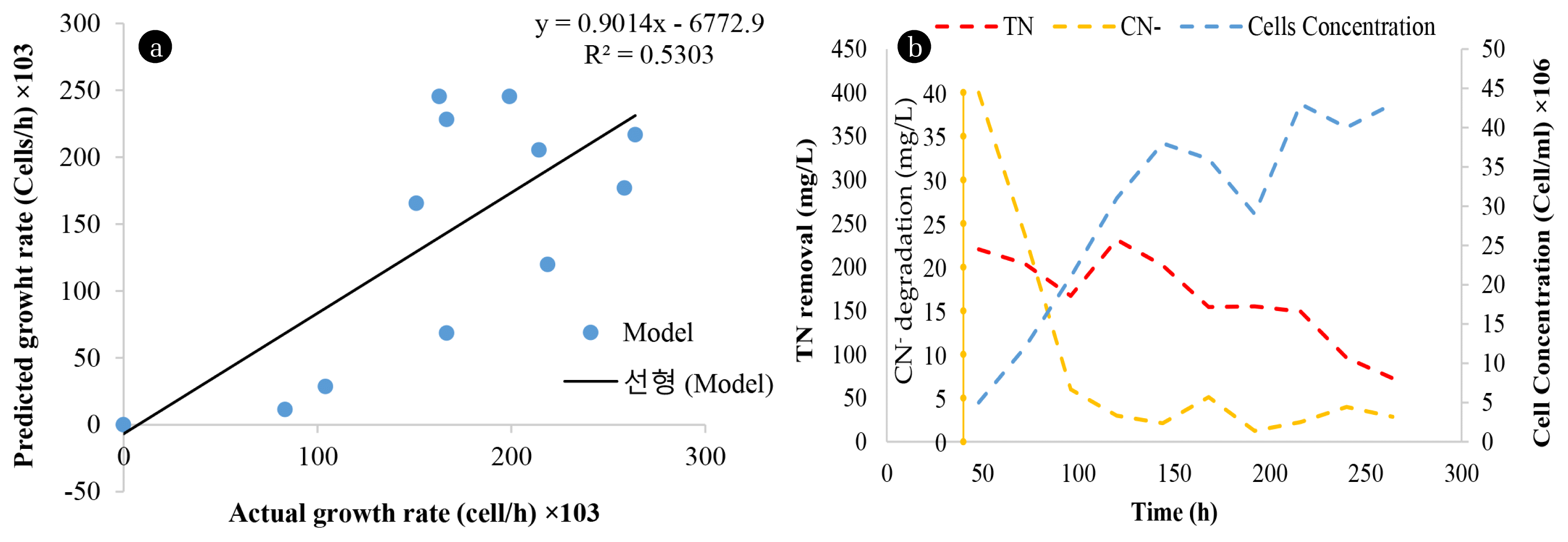
Fig. 5A comparison of the prediction ability of RSM and cybernetic models. (a) prediction of TN removal efficiency by RSM model; (b) parity plot comparing predicted total nitrogen removal efficiency and actual total nitrogen removal efficiency by RSM; (c) Rate of TN removal predicted by cybernetic model; (d) parity plot for comparing the predicted rate of TN removal with the actual rate of TN removal by cybernetic model. 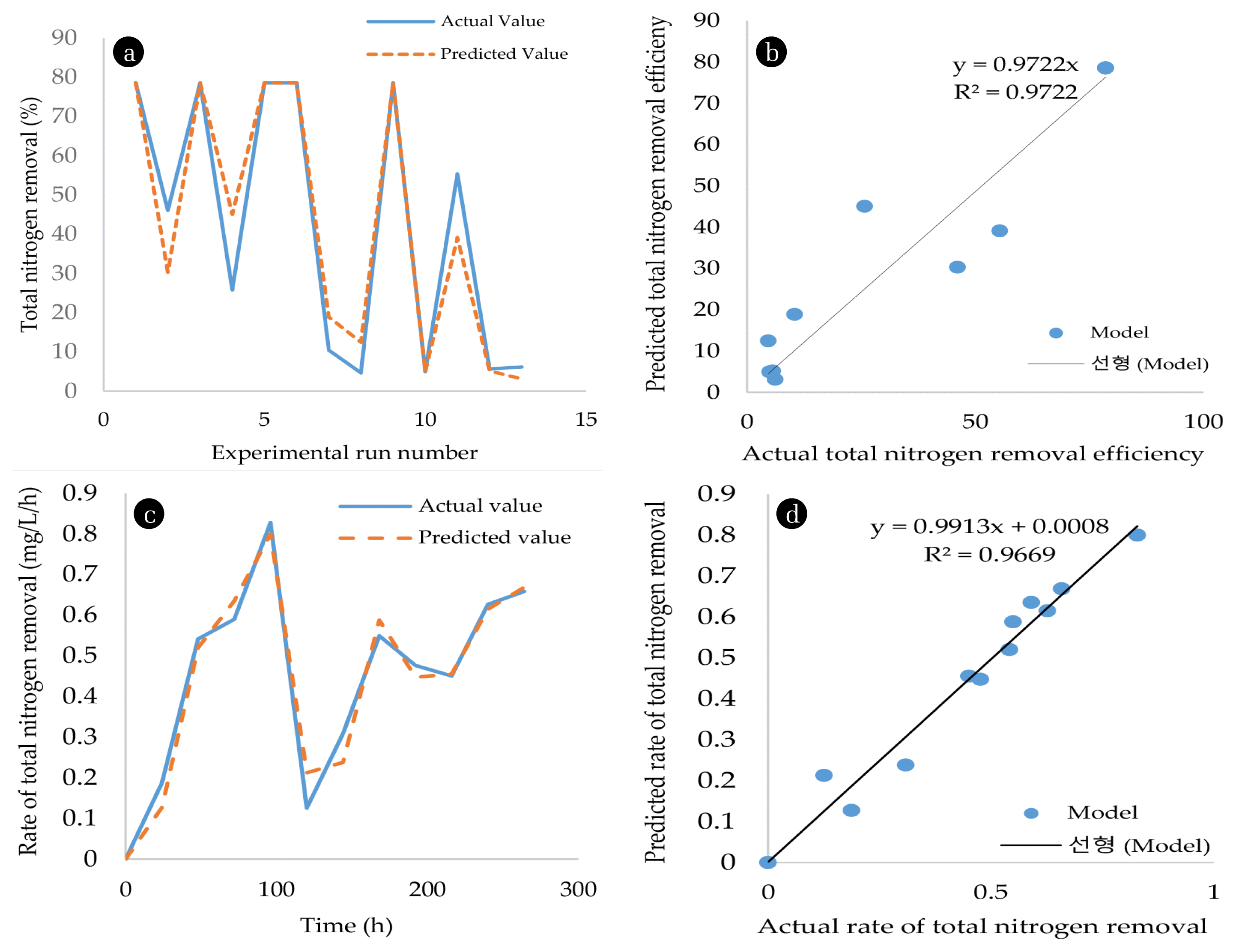
Fig. 6(a) NH4-N and CN- removal/ degradation. (b) NO2-N, and NO3-N removal simulation of the cybernetic model. (c) level of key enzyme e1 over time. (d) level of key enzyme e2 over time. (e) level of key enzyme e3 over time. (f) level of key enzyme e4. 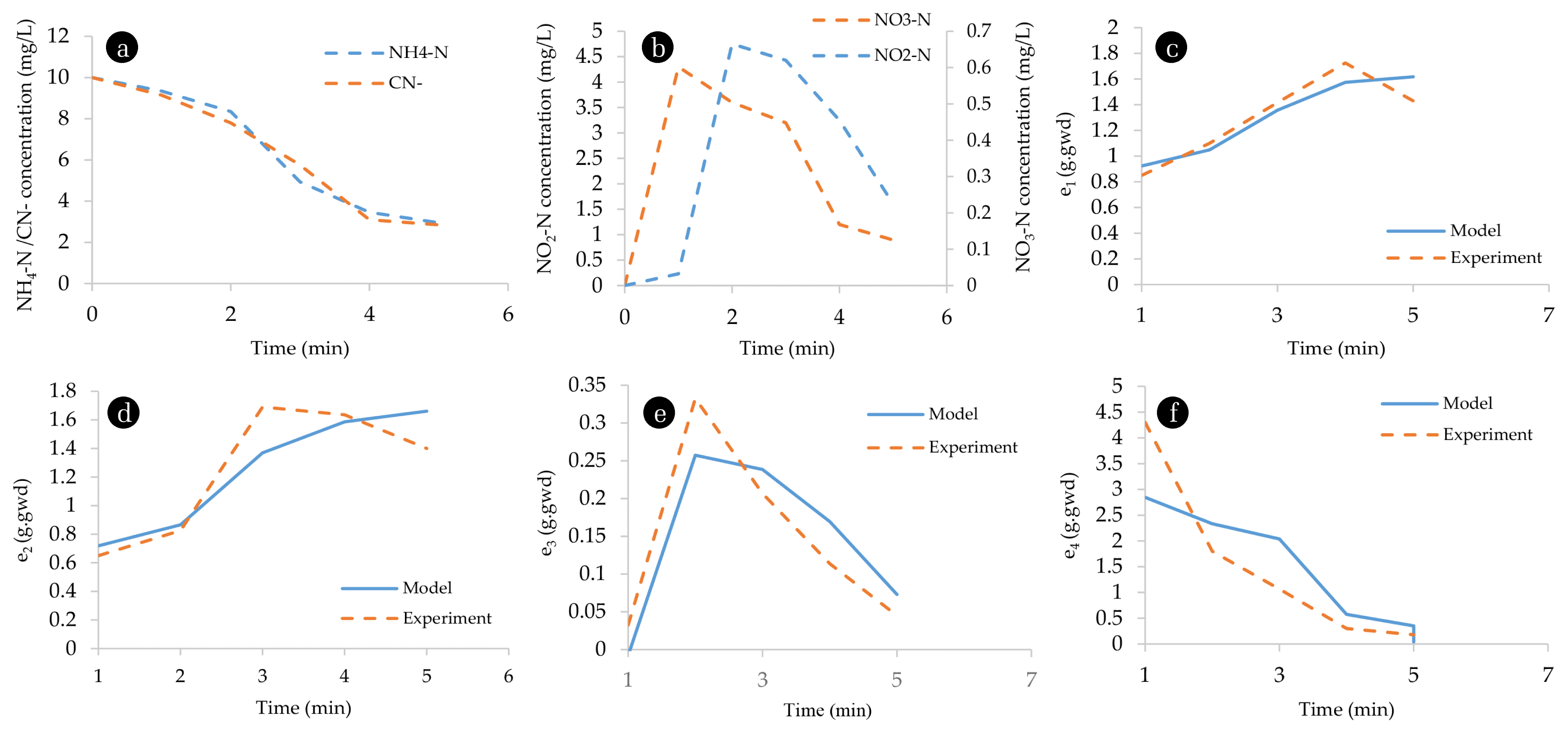
Table 1Analysis of Variance (ANOVA) of the Quadratic Parameters for SNaD Process Used for TN Removal under CN- Conditions Table 2Model Parameter Estimations |
|
||||||||||||||||||||||||||||||||||||